**This post is a part of the series on the 2019 AGA Presidential Symposium – Sex and Asex: the genetics of complex life cycles**

About the author: Dr. Stacy Krueger-Hadfield is an evolutionary ecologist and assistant professor of biology at the University of Alabama at Birmingham. She works on life cycle and reproductive system evolution in algae and invertebrates. She is the Editor of Social Media and a member of the AGA Council. Follow Stacy on Twitter @quooddy or via her lab’s website.
Concerning haplodiplontic population genetics
The story for the two companion papers – one led by Solenn Stoeckel (Stoeckel et al. 2021) and one led by myself (Krueger-Hadfield et al. 2021) – began in late 2010 or early 2011. As part of the intro to my dissertation, I was compiling population genetic data that had been generated in haplodiplontic algae – with the important caveat that both haploid and diploid stages had been sampled.
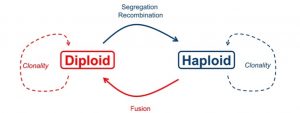
While all sexual taxa undergo an alteration of haploid and diploid stages, in the majority of macroalgae (= seaweeds; a polyphyletic group including lineages in the Stramenopiles [browns, like kelps] and the Archaeplastids [greens, like sea lettuce; and reds, like Irish moss) have free-living, wholly distinct haploid and diploid adults (Figure 1). In other words, the haploid stage isn’t just a single-celled, short-lived gamete. Instead, it’s as if the gametes grow up into adults.
My literature search didn’t take long – there were only a handful of studies that had genotyped haploid gametophytes and diploid sporophytes.
I took another tack. I contrasted data generated by Carolyn Engel during her PhD on Gracilaria gracilis with data I’d been generating on Chondrus crispus. We had two taxa that occupied similar intertidal habitats along the coast of France, but had totally different mating systems. Gracilaria male and female gametophytic mates were unrelated. Chondrus male and female gametophytes, on the other hand, were quite happy to mate with full sibs on a regular basis. Yes, these two species are phylogenetically distinct and they have different distribution patterns (Figure 2), but it was curious that they were so different from one another in terms of their mating system. Red algae are united by some really weird quirks – zygotic amplification, no motile propagules, etc. Yet, with 1 outcrosser and 1 seller, it’s kinda hard to determine whether red seaweeds, in particular, tend to being outcrossers or selfers (it turns out that even if you have separate males and females, selfing is still possible in these taxa).
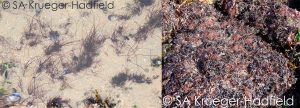

These musings led to discussions with my PhD advisor, Myriam Valero, about a review on seaweed reproductive systems – highlighting the lack of data – but the idea languished in the rush to finish my PhD and starting a post-doc in the UK.
The post-doc goes south
Not south as in a bad way. Literally south. I moved from the UK to South Carolina. A chance meeting with Sean Hoban led to a collaboration on a paper providing some guidelines for sampling haplodiplontic taxa (Krueger-Hadfield and Hoban 2016). Our ideas and protocols came just in time for our Northern Hemisphere-wide sampling effort of an invasive red seaweed. One stop on this summer extravaganza was back to my old stomping grounds in France.
Marie-Laure Guillemin was in Roscoff at the same time. I still have the notes somewhere, but I am sure you have figured out by now, that the paper was to remain a figment of our imaginations for a bit longer.
A newPI journey (sometimes) in the dark
In 2016, I started my lab at UAB and the mating system paper flitted around just out of reach while I was trying figure out how to run the lab, recruit people, teach, and do research all at the same time.
Ideas about haplodiplontic population genetics nagged me. I was unsatisfied with ways we had to collect and analyze data in haplodiplontic taxa, but no real direction on how to rectify those problems. And, so the paper waited.
Edgefield
We stumbled along to 2019 when Maria Orive was putting together the Sex/Asex symposium that would be at the Edgefield hotel outside Portland, OR. I will admt that I blanched when I saw I’d be speaking third, after Sally Otto and Laura Katz. Not only speaking after them, but then also participating in a panel with them.
Solenn Stoeckel had also been invited to speak and he and I had mused over the haplodiplontic pop gen conundrum earlier in 2019 without gaining much momentum. We were also now all part of the ANR-funded CLONIX-2D consortium (led by Solenn) and were busy trying to figure out the best ways to assess partial clonality across a range of taxa.
Off the cuff, I told Solenn about the work I’d done with Sean and how I wanted to figure out better ways to analyze haplodiplontic taxa. The whole diploid data set and a haploid data set seemed artificial when the ploidy stages are exchanging genes! We started sketching out the ideas to simulate haplodiplontic populations and then compare those theoretical predictions to what has actually been shown in nature. We finally were amalgamating dormant ideas …
If you recall, the summer of 2019 was a VERY hot summer in Europe. Solenn had to pause our simulations when the cluster in Rennes was overheating. It was too hot to run our simulations! Was this a harbinger of what was to come in 2020? At the time, we were blissfully unaware.
Meanwhile, Myriam, Christophe, Marie-Laure, and I set about finding data sets in mosses, ferns, or algae where both the haploids and diploids were genotyped. We came up with a list made only slightly longer than the one I’d compiled almost 10 years earlier – slightly longer as we’d added some mosses and included a lot of our own work! There weren’t any ferns, but not unlike kelps, one stage is microscopic so not really very easy logistically for genotyping.
Farewell to normal life
2020 dawned in a frantic rush to finish both the ‘theoretical’ paper led by Solenn and the ’empirical’ paper led by me. Inevitably, we had to ask for an extension. And, then, dear reader, we all know what happened come March …
While the theoretical paper was submitted fashionably late, the empirical paper lingered … as many New PI’s are acutely aware, all our hard work getting our labs up and running vanished into the ether. No one was allowed in the labs and teaching moved online. There was a never ending round of squelching fires just in time for another one to erupt.
There were moments in early lockdown when I had incredibly proud moments – like making this figure:
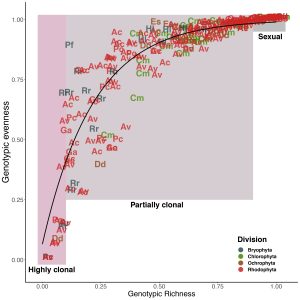
Yet, every time I returned to to the empirical paper, my attention would be yanked in another direction. Finally, on 1 November 2020, I’m slightly ashamed to admit, we submitted the ’empirical paper’ – a full nine months to the day from when it had been due. I can’t thank Maria and Anjanette Baker enough for being patient with us as we kept asking for extension after extension after extension.
Epilogue
In editing the blog posts for the special issue, I was struck by the tales each author told about the long journey to publication or feelings of ‘not belonging’ – both of these papers encapsulate those sentiments for me. I’m not a mathematical wiz, yet, I’ve been part of some rather complex papers and learned a ton in the process. I also struggled with the delays that kept pushing the ’empirical’ paper down the triage list in deference to other things that warranted my undivided attention.
While the theoretical (Stoeckel et al. 2021) and empirical papers (Krueger-Hadfield et al. 2021) aren’t what Myriam and I had envisioned way back when, the unexpected journey and percolation of those ideas for nigh on a decade led to two papers of which I am immensely proud. We have our work cut out for us though – there is still lots to do in terms of generating data in haplodiplontic taxa as well as writing another paper or two that have sprouted as a result of this work.
And, I promised Anjanette, I’d never have a paper in so late again.
References



