
About the Blog Author: Jeremy is an evolutionary ecologist and NSF postdoctoral fellow working with Catherine Linnen at the University of Kentucky Department of Biology. He is broadly interested in adaptation, speciation, population genomics and chemical ecology of insects. Follow him on Twitter @flydrocarbon.
Biological introductions can be seen as “natural experiments” that provide unique insights into evolutionary processes. Invasive plant-feeding (phytophagous) insects are of particular interest for studying adaptation, as introductions frequently require rapid colonization of new host. However, invasions are often limited by reduced genetic diversity, leading to a quandary known as the “genetic paradox of invasions” (Frankham 2005). A common “solution” to this paradox is that many introductions involve multiple invasive waves that introduce additional genetic diversity required for overcoming adaptive challenges (Rius and Darling 2014). Therefore, detailing the history of invasion on a demographic and genetic level is crucial for understanding how species adapt to novel environments.
In my EECG Embarkation post, I introduced my study organism, Diprion similis, a European pine sawfly that invaded North America in 1914. Upon introduction, this species colonized eastern white pine (Pinus strobus), which is avoided by most native pine sawflies because its thin needles pose a challenge for successful oviposition (Bendall et al. 2017). I was honored to be awarded an EECG grant in 2021 to investigate if multiple invasive waves were responsible for successful colonization of a novel host in this species and provide further insight into overcoming adaptive challenges during biological introductions.
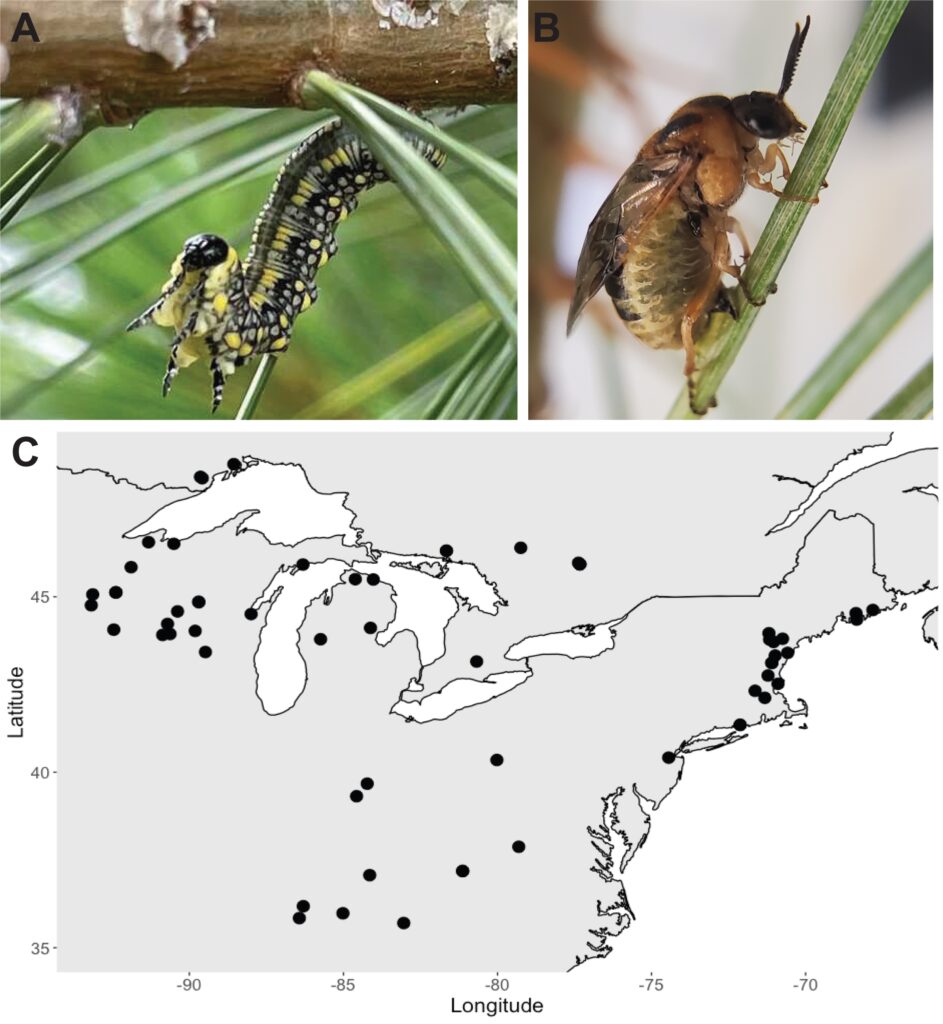
I’m excited to share the results of this project now published in the Journal of Heredity. First, we collaborated with an excellent team from the USDA-ARS (Sheina Sim, Scott Geib, Brian Sheffler) to produce a reference genome for this species. This resulted in a high-quality reference that will be a powerful genomic resource for using D. similis as an emerging model for genomic study of biological introductions. This genome was then put to use to map low-coverage whole-genome sequence data from 64 D. similis females sampled from across eastern North America to use for population genomic analyses (Fig 1). The ultimate goal was to use geographically diverse individuals to sample genomic diversity in the introduced range and evaluate evidence for multiple introductions.
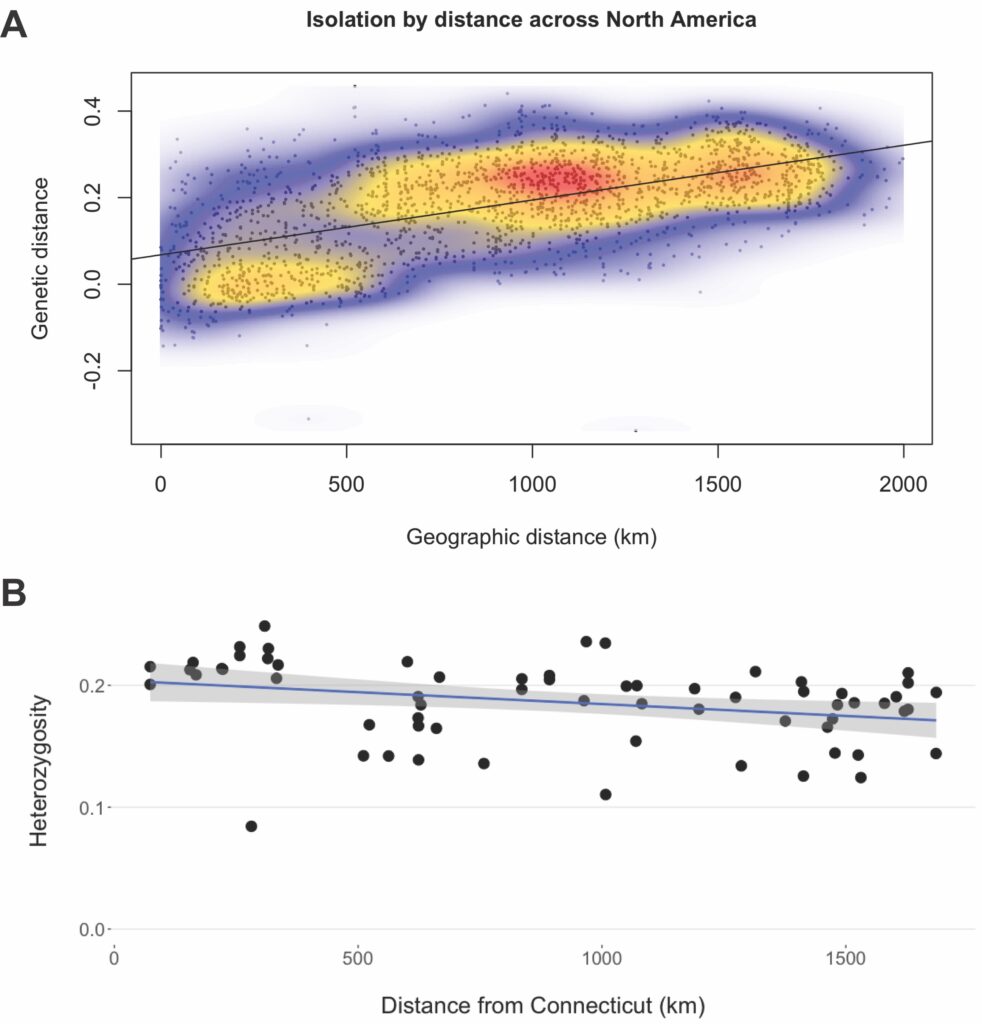
Our results indicate good evidence that D. similis was introduced in a single invasive wave, contrary to our expectations and those most commonly seen in invasive biology literature (Rius and Darling 2014). The evidence for this single introduction is three-fold, based on expectations under single or multiple introduction scenarios. First, a multiple-introduction scenario from genetically distinct groups is expected to be detectable as genetically distinct clusters. However, in our data, both model-based and model-free clustering analyses reveal a single, continent-wide population of D. similis, suggesting either a single invasion or multiple waves of genetically similar individuals. Second, assuming sufficient time to reach drift-migration equilibrium, a single introduction scenario is expected to show a mostly continuous pattern of isolation-by-distance (IBD). Our results from D. similis match this expectation, with a continuous spatial pattern to genetic diversity in the introduced range (Fig 2A). Lastly, in a single introduction, genetic diversity is expected to decrease monotonically from the original point of introduction as populations are smaller on the edges of range expansion. Similar to our IBD result, we found that heterozygosity was negatively correlated with distance from Connecticut, the well-established point of introduction (Fig 2B). Together, these lines of evidence paint a strong picture that D. similis likely invaded in a single major wave.
Because our results indicate a single introduction event, the genetic paradox is still not “solved” in this case and begs the question: “how did D. similis successfully colonize a new host given the limited genetic diversity of its introduction?”. Some possibilities—in brief—are: 1) unlike other pine sawflies, D. similis may not face an adaptive challenge from P. strobus’ thin needles. 2) multiple introductions have occurred but using only modern samples might hide a history of additional invasive waves that either a) were outcompeted and went extinct, or b) thoroughly admixed far enough in the past the signature is lost. These possibilities open new research avenues that I am exploring using museum specimens as part of a larger NSF funded project. Overall, our findings and implications for future work in this system generate an interesting and nuanced conversation, and I implore curious readers to read about it at length in our newly published article.
I am grateful to the AGA, the Journal of Heredity, and my co-authors for making this project and the resulting manuscript possible. This work lays the groundwork for many exciting research directions using D. similis as a study system, and funding provided by this EECG award has gone a long way to generate genomic resources and data on demography and population dynamics in a non-model system.
References:
Bendall, E. E., Vertacnik, K. L., & Linnen, C. R. (2017). Oviposition traits generate extrinsic postzygotic isolation between two pine sawfly species. BMC Evolutionary Biology, 17(1), 115.
Frankham, R. (2005). Resolving the genetic paradox in invasive species. Heredity, 94(4), 385.
Rius M, Darling JA (2014) How important is intraspecific genetic admixture to the success of colonising populations? Trends in Ecology & Evolution, 29, 233–242.



