
About the author: Dr Helen Taylor is a conservation geneticist who studied for her PhD in New Zealand, working on inbreeding in little spotted kiwi. She went on to undertake postdoctoral research on inbreeding and male fertility in passerines and, at that point, became interested in the integration of genetics into conservation management. After eight years in New Zealand, Dr Taylor left academia and headed back to the UK to work as conservation programme manager at the Royal Zoological Society of Scotland. Find out more here: www.helentaylorscience.com

In this series, written specially for the AGA Blog, Dr. Taylor will be exploring the gap between conservation genetics research and conservation implementation, showcasing some examples of how the gap is being closed for various species and projects, and exploring what it means to be a conservation geneticist in the modern sense (aka, why at least some of this is our fault and we need to do better). Strap in for a rollercoaster ride through the politics of conservation genetics, viewed through the lens of a former academic who now works in conservation management.
The story of Tasmanian devil conservation reads like a page-turning thriller. A race against time to rescue Australia’s last surviving large marsupial carnivore from a deadly transmissible facial cancer as the disease sweeps across the last refuge of a fierce yet vulnerable species. This tale has it all, including (best of all!) conservation genetics! Tasmanian devil conservation is, happily, also a story of genetics informing conservation management in a way that, as we’ve seen in my previous posts, is often sadly lacking. Here, I take a look at what makes this conservation genetics project stand out.

Doomed devils?
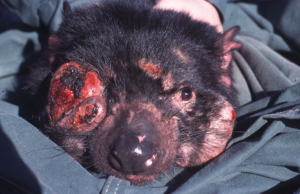
Tasmanian devils have not had a great run over the past few thousand years. Their name is misleading; this marsupial was once widespread through mainland Australia, but a hypothesised combination of climate change, dingoes, and human intensification1 saw them isolated on Tasmania by around 3000 years ago2. Thanks to this drastic population decline, and further declines on Tasmania, devils have relatively low genetic diversity, including at immune loci3. This lack of genetic diversity makes them vulnerable to chance environmental challenges. Particularly challenges like Devil Facial Tumour Disease (DFTD), a gruesome and deadly transmissible cancer that has reduced Tasmanian devil numbers by almost 80% across their range since the late 90s4.
Devil Tools and Tech: an origins story
In 2006, as DFTD continued to ravage the devils of Tasmania, the race was on to gather up disease-free individuals to form an insurance population. However, with founders all having been caught in a relatively small geographical area ahead of the disease front, there were concerns regarding genetic diversity and inbreeding in this captive “clean” population of devils5. There was a clear need for careful genetic management of the insurance population, but low genetic diversity in devils meant that tools like microsatellite markers lacked the statistical power to estimate relatedness between devils5.
The Devil Tools and Tech programme was set up with the intention of developing genetic tools to assist the conservation management of Tasmanian devils6. The programme’s leaders made their goal of bridging the conservation genetics gap explicit in the way they set up the project. In their words, they wanted to take genetic tools “from the lab bench to the forest floor”. This was (and still is) a genetic and genomic research programme driven by conservation management needs. Additionally, and crucially, the programme bought together teams from government, industry, and academia.
A conservation genetics collaboration
The partnership for Devil Tools and Tech reads like a conservation chain reaction. The government-run Save the Tasmanian Devil initiative needed help managing an insurance population, so they got the Zoo and Aquarium Association of Australia involved. The Zoo folks saw the need for genetic management, and invited conservation genetics researchers at the University of Sydney to come on board. Having found the skills, the team still needed to pay the bills, and so San Diego Zoo Global came to the party and funded a postdoctoral position on the project.
Since coming together as a team, the various people involved in Devil Tools and Tech have produced over 40 research publications, all with implications for devil conservation management. From designing reference-based SNP calling pipelines for non-model species7, to refining captive breeding to avoid declines in reproductive success8 (in our very own Journal of Heredity, WOOT!) the programme has addressed multiple questions that matter to Tassie devil conservation, and that apply to the conservation of many other species too.

A bridge to success
There are some key points about Devil Tools and Tech that have made it so successful in getting genetic data integrated into the management of a threatened species. Hopefully, you’ll spot some themes from my previous post.
First, the research programme was guided by clearly stated conservation needs rather than being driven by “publish or perish” pressure. By employing cutting-edge genomic technology to meet those needs where possible, the Tools and Tech team managed to generate a large volume of highly publishable work, while enhancing the conservation management of Tasmanian devils. Bridging the conservation genetics gap does not have to mean sacrificing high-impact science.
Additionally, the organisations involved in the programme understood that each of them alone could not save Tasmanian devils – they had to work together as a collective to achieve this. The combination of participants from government, industry, and academia made it far more likely that the data produced would be used to make recommendations that would be integrated into management and policy.
Managing multi-faceted partnerships like the Devil Tools and Tech programme is no mean feat. All the organisations involved invested both intellectually and financially in the project, meaning that everyone had more skin in the game than just their desire to see devils conserved. Regular communication between project partners was key, as was allowing managers to have early access to data ahead of publication. Waiting for peer review sucks at the best of times, but it’s even worse when you’re waiting for recommendations on how to prevent the eventual extinction of the species you manage.
Continuing to evolve
By employing all the tactics and techniques described above, the Devil Tools and Tech project has become a real force for proactive conservation genetic management. The project continues to evolve to address new questions regarding the importance of the microbiome in the post-release phase of captive-bred animals9, and using poo to assess dietary needs and seek out much needed additional genetic diversity in remote, potentially untapped, populations of devils10.
Finally, if you’re a fan of STEMINISM (and, if you’re not, maybe go and have a word with yourself), then don’t just take my word for it that this is a great project. Go read the stories of some of the women who made this project such a success11. And then come back here in time for my next conservation genetics gap blog post!
References
- Brown, O. J. F. Tasmanian devil (Sarcophilus harrisii) extinction on the Australian mainland in the mid-Holocene: multicausality and ENSO intensification. Alcheringa: An Australasian Journal of Palaeontology 30, 49–57 (2006).
- White, L. C., Saltré, F., Bradshaw, C. J. A. & Austin, J. J. High-quality fossil dates support a synchronous, Late Holocene extinction of devils and thylacines in mainland Australia. Biology Letters 14, 20170642 (2018).
- Morris, K., Austin, J. J. & Belov, K. Low major histocompatibility complex diversity in the Tasmanian devil predates European settlement and may explain susceptibility to disease epidemics. Biology Letters 9, 20120900 (2013).
- Hawkins, C. E. et al.Emerging disease and population decline of an island endemic, the Tasmanian devil Sarcophilus harrisii. Biological Conservation 131, 307–324 (2006).
- Hogg, C. J. et al.Influence of genetic provenance and birth origin on productivity of the Tasmanian devil insurance population. Conservation Genetics 16, 1465–1473 (2015).
- Hogg, C. J. et al.“Devil Tools & Tech”: A Synergy of Conservation Research and Management Practice. Conservation Letters 10, 133–138 (2017).
- Wright, B. et al.From reference genomes to population genomics: comparing three reference-aligned reduced-representation sequencing pipelines in two wildlife species. BMC Genomics 20, 453 (2019).
- Farquharson, K. A., Hogg, C. J. & Grueber, C. E. Pedigree analysis reveals a generational decline in reproductive success of captive Tasmanian devil (Sarcophilus harrisii): implications for captive management of threatened species. Journal of Heredity 108, 488–495 (2017).
- Chong, R. et al.Looking like the locals – gut microbiome changes post-release in an endangered species. Animal Microbiome 1, 8 (2019).
- Grueber, C. E. et al.Genetic analysis of scat samples to inform conservation of the Tasmanian devil. Australian Zoologist 40, 492–504 (2020).
- Fox, S., Hogg, C. J., Grueber, C. E. & Belov, K. Devil women. Pacific Conservation Biology 24, 271–279 (2018).
- Grueber, C. E. et al.Increasing generations in captivity is associated with increased vulnerability of Tasmanian devils to vehicle strike following release to the wild. Scientific Reports 7, 2161 (2017).



