Connecting genotypes to the complex phenotypes they produce is a fundamental goal of genetics. Variation in coat color pigmentation, partly due to the relative ease at which different patterns can be identified, is one of the best-characterized traits at the genetic level, with examples from sheep (Zhang et al., 2017), cattle (Li et al., 2016), and dogs (Newton et al., 2000), among other animals. While there are still plenty of unknowns, one reoccurring theme in this line of research is that similar phenotypes are not necessarily underlain by the same genetic variants. Instead, these convergent phenotypes tend to result from different mutations in the same gene or mutations in different genes that are part of the same molecular pathway that ultimately produces the phenotype. In the case of the splashed white phenotype in horses, which is the proper term for “looks like the legs and belly were dipped in white paint,” both of these molecular mechanisms of convergence had been previously documented (Hauswirth et al., 2012). Five variants- three in the melanocyte inducing transcript factor MITF, and two in PAX3, a known transcriptional activator of MITF– had been previously associated with the splashed white phenotype in horses (Hauswirth et al., 2012). These mutations range from relatively small insertions (10 bp, SW1) and deletions (5 bp, SW3), to large deletions (64 kb, SW5) and single base-pair nonsynonymous mutations (SW2 & SW4). Moreover, mutations in EDNRBand KIT have been associated with other white patterned phenotypes in horses (e.g., Brooks and Bailey, 2005; Haase et al., 2007).
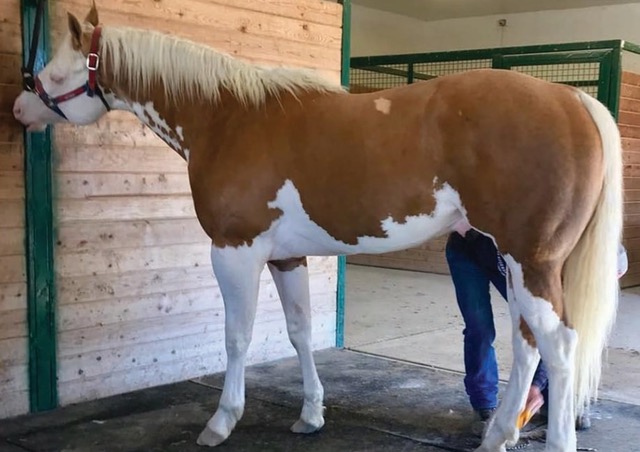
Magdesian, et al., (2020) recently investigated a phenotypic puzzle that most wildlife geneticists can only dream about: a splashed white stallion (Figure 1) with non-splashed white parents that did not carry any of the previously characterized mutations associated with the phenotype. Hypothesizing that an uncharacterized de novomutation underlay the splashed white phenotype, the authors used full genome sequencing to investigate.
The authors began their search with the genomic regions that had been previously identified to carry mutations that impact the white phenotype, namely, MITF, PAX3, EDNRB, and KIT among a few others. After eliminating two single base-pair mutations in EDNRBand KIT that were not predicted to have any appreciable effect on protein structure, the authors searched for structural variants. Using the sequence data visualization tool Integrative Genomics Viewer (IGV) (Robinson et al., 2011), the authors identified an 8710 bp deletion in MITF. Following confirmation with Sanger sequencing, the stallion in question was determined to be heterozygous for the mutation, while neither the sire nor the dame carried a copy. The authors then used the program PredictSNP (Bendl et al., 2016) to evaluate how this deletion would influence the final protein product, and found that it induced a premature stop codon, resulting in a truncated helix-loop-helix DNA binding domain. Lastly, Magdesian et al. genotyped the stallion’s 11 offspring, four of which were splashed white and one of which was all-white, for the deletion. All five of the white offspring were found to be heterozygous for the novel MITFmutation, while the all-white offspring also carried the SW1 allele and a previously characterized dominant white allele (W20).
Taken together, and in the absence of any other novel mutations in splashed white associated genes, the authors concluded that this 8710 bp deletion in MITF is likely to be the causal mutation of the splashed white phenotype in the focal stallion and the splashed white offspring, with a possible compounding effect of carrying multiple white patterning genes, as demonstrated by the all-white offspring. While this latter observation could be important for targeted horse breeding efforts, the authors suggest future research on the MITFprotein structure and function to confirm the overall impact on the splashed white phenotype in horses.
References

Elizabeth Heppenheimer received her PhD in 2019 in Ecology and Evolutionary Biology from Princeton University where she worked on canid genetics. She currently works in academic publishing and science communication. Follow her @LizHeppenheimer