
About the author: Dr Helen Taylor is a conservation geneticist who studied for her PhD in New Zealand, working on inbreeding in little spotted kiwi. She went on to undertake postdoctoral research on inbreeding and male fertility in passerines and, at that point, became interested in the integration of genetics into conservation management. After eight years in New Zealand, Dr Taylor left academia and headed back to the UK to work as conservation programme manager at the Royal Zoological Society of Scotland. Find out more here: www.helentaylorscience.com
In this series, written specially for the AGA Blog, Dr. Taylor will be exploring the gap between conservation genetics research and conservation implementation, showcasing some examples of how the gap is being closed for various species and projects, and exploring what it means to be a conservation geneticist in the modern sense (aka, why at least some of this is our fault and we need to do better). Strap in for a rollercoaster ride through the politics of conservation genetics, viewed through the lens of a former academic who now works in conservation management.
Wildcats are cute. Let’s get that out of the way first. Their kittens are playful, blue-eyed floofsters that capture the hearts of almost everyone who sees them. In Britain, wildcats are also Critically Endangered. Thanks to persecution and habitat loss, the British population of this species is now restricted to the highlands of Scotland, where it has become so rare that a new threat has emerged; hybridisation with domestic and feral cats.

Despite 1.1 million years of separation1, wildcats and their domesticated cousins readily interbreed. Such interspecies mingling increases disease transmission into the wildcat population2 and produces hybrid descendants that can be extremely difficult to tell apart from genuine wildcats3. This is a problem. It’s a problem if you want to trap and neuter non-wildcats to mitigate the threat, and it’s a problem if you want to select genuine wildcats to breed for release.
Fortunately, a team of canny conservation geneticists found a solution. It’s a solution that shows the value of integrating conservation genetics into species management planning, and clearly illustrates how zoo-based conservation teams are well-equipped to bridge the conservation genetics gap.
We’re going to the zoo, zoo, zoo…
The Royal Zoological Society of Scotland (RZSS) is a wildlife conservation charity that runs two zoos (Edinburgh Zoo and Highland Wildlife Park) and (full disclosure) is also my current employer. Hidden away behind the rhino house at Edinburgh Zoo is one of RZSS’ most valuable assets to conservation, the WildGenes laboratory – the UK’s only zoo-based conservation genetics lab.

The RZSS WildGenes team provides conservation genetics support to a wide variety of conservation projects, from illegal wildlife trade tracking in Cambodia, to non-invasive monitoring of capercaillie in Scotland, and population management of dama gazelle in North Africa. The team has a strong track record of integrating conservation genetics into management plans. More importantly for this particular blog post, the team also developed a genetic test for hybridisation in wildcats4.
Genetic detective work
The wildcat hybrid test is a faster, more-cost effective version of a Swiss test for similar purposes5–7. The Swiss test used 83 SNP makers, while the WildGenes test employs 35 SNPs that distinguish wildcats from domestic and feral cats. They do so with lower confidence than the original 83 SNP test but, given the common financial limitations faced by most conservation projects, it’s a worthwhile trade-off4.
Based on this test, the WildGenes team produced a threshold to determine when a cat was ‘good enough’ to be classed as a wildcat. This threshold was a balance between inadvertently losing good wildcat DNA from the population and reducing introgression. The cut off is currently set at >75% wildcat DNA=good enough4. Testing shows that the 35 SNP markers selected for the test perform almost as well at detecting hybrid cats as a panel of over 3,000 SNPs produced via ddRAD3. Developing this test, however, was just part of a fully integrated approach to wildcat conservation in Scotland.
Just how wild are those wildcats?
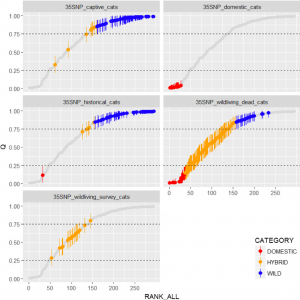
How many genuine wildcats remain in Britain? There are effectively two populations of wildcats in the country – those scattered across the highlands of Scotland, and those in the captive breeding programme across more than 30 British zoos,wildlife parks and private holders. The WildGenes team tested Scottish cats captured in the wild as part of the trap-neuter-release programme, and dead wildcats collected mainly as roadkill, plus cats from across the captive population. They compared the genetic test scores of these animals with those from known domestic cats, and historical samples of free-living wildcats from Scotland3.
The results of the study underlined how big an issue hybridisation is for wildcats in Scotland. All “wildcats” tested from live capture scored as hybrids (two genuine wildcats have been found living wild since the study), as did the vast majority of roadkill “wildcats”. This was very different to the historic situation, with most museum specimens testing as genuine wildcats. So far, so gloomy. However, the majority of cats in the captive breeding programme tested as genuine wildcats3. Huzzah! There was now a reliable way of identifying valuable genuine wildcats, and a good source population for the species in Britain.
Thinking big and bridging all the gaps
At this point, WildGenes being part of a zoological organisation really comes into its own. The current plans for wildcat conservation in Scotland hinge on breeding wildcats to release back into the Highlands to boost the numbers so that hybridisation is less of a threat. In addition to the WildGenes team, RZSS is home to the studbook keeper for the wildcat breeding programme, a field conservation unit, and a host of experienced vets and animal managers, putting the organisation in a great position to lead an integrated wildcat conservation programme.
RZSS is now the home of the Saving Wildcats project, an initiative that brings together RZSS, NatureScot (the main government conservation agency), Forestry and Land Scotland, the Cairngorms National Park Authority, Nordens Arkzoo in Sweden, and Junta de Andaluciain Spain. RZSS’ Highland Wildlife Park site is about to become home to a large breeding-for-release facility (one of the first of its kind in the UK), where wildcats selected from the captive population using the 35-SNP test will be bred with minimal human interaction, ready for release into a specially designated area in the Cairngorms. The project is funded by a large EU LIFE grant, running over six years.

The conservation genetics for wildcat management is also developing rapidly. Further research using thousands of markers discovered via ddRAD and whole genome sequencing is underway in collaboration with the University of Bristol to establish a deeper understanding of hybridisation, inbreeding and relatedness in the species. DNA metabarcoding tools will also be used to monitor the diet of the released cats.
Saving Wildcats brings together RZSS’ expertise, and teams it with the complementary skills and expertise of the other project partners. The project also involves social science, science communication, habitat management and rigorous monitoring. It is a brilliant showcase for how zoo conservation science teams are ideally placed to bring conservation genetics to the heart of conservation management, and bridge multiple strands of conservation management.
Zoo conservation genetics teams in the spotlight
RZSS is not alone. San Diego Zoo Global, the Chicago Zoological Society,The Smithsonian’s National Zoo, Omaha’s Henry Doorly Zoo, Copenhagen Zoo, The South African National Biodiversity Institute, and the Antwerp Zoo Society all have their own laboratories and conservation genetics staff and all work to integrate genetics into species conservation management.
If you’re a conservation genetics graduate looking for a position where your skills can be applied to conservation programmes in a meaningful way, then breaking out of academia and into zoo conservation is definitely worth considering. The story of the wildcat in Scotland is a perfect example of why.
References
- Li, G., Davis, B. W., Eizirik, E. & Murphy, W. J. Phylogenomic evidence for ancient hybridization in the genomes of living cats (Felidae). Genome research26, 1–11 (2016).
- Kilshaw, K. Scottish wildcats. (2011).
- Senn, H. v et al.Distinguishing the victim from the threat: SNP-based methods reveal the extent of introgressive hybridization between wildcats and domestic cats in Scotland and inform future in situ and ex situ management options for species restoration. Evolutionary Applications12, 399–414 (2019).
- Senn, H. & Ogden, R. Wildcat Hybrid Scoring For Conservation Breeding under the Scottish Wildcat Conservation Action Plan. (2015).
- Nussberger, B., Greminger, M. P., Grossen, C., Keller, L. F. & Wandeler, P. Development of SNP markers identifying European wildcats, domestic cats, and their admixed progeny. Molecular Ecology Resources13, 447–460 (2013).
- Nussberger, B., Wandeler, P. & Camenisch, G. A SNP chip to detect introgression in wildcats allows accurate genotyping of single hairs. European Journal of Wildlife Research60, 405–410 (2014).
- Nussberger, B., Wandeler, P., Weber, D. & Keller, L. F. Monitoring introgression in European wildcats in the Swiss Jura. Conservation Genetics15, 1219–1230 (2014).



