**The AGA grants EECG Research Awards each year to graduate students and post-doctoral researchers who are at a critical point in their research, where additional funds would allow them to conclude their research project and prepare it for publication. EECG awardees also get the opportunity to hone their science communication and write posts over their grant tenure for the AGA Blog. In the first in the series, our EECG awardees write about their research and their interests as an ’embarkation’.**

About the Blog Author: Robert Driver is a PhD candidate in the Department of Biology at East Carolina University. He researches the molecular evolution of gene families involved in the sensory perception of birds. Follow Robert on Twitter @rdriver215.
Olfaction plays a critical role in animal foraging, predation, recognition, and territorial behavior (Touhara and Vosshall 2009). Vertebrates detect odor molecules with olfactory receptors (ORs), a gene family expressed in the olfactory sensory neurons of the olfactory epithelium (OE; Buck and Axel 1991, Strotmann et al. 1992). An incredible variety of chemicals are perceived as smell by animals (Saito et al. 2009). To bind this range of chemicals, ORs constitute the largest gene family in vertebrates, with nearly 2,000 genes in some mammals (Niimura et al. 2014). The number of ORs in a species’ genome can be used to derive total genomic repertoire counts (Niimura et al. 2014), but not all of the genomic repertoire will be functional or relevant to the olfactory system. Many ORs are expressed in tissues outside the olfactory system (Maßberg and Hatt 2018). Within this context of a complex gene family, understanding the function (or lack thereof) of specific ORs is a major challenge. Gene expression studies of the OE can distinguish ORs that likely bind odorants from ORs with other physiological roles and non-functional pseudogenes. Expression studies of the OE have occurred in all vertebrate classes (e.g., Marchand et al. 2004, Kolmakov et al. 2008), but have not occurred in birds until recently, with only one study targeting a single species (Sin et al. 2022).
Birds are the most speciose class of terrestrial vertebrates, inhabiting nearly all land environments and with diverse breeding and foraging habits (Jetz et al. 2012). I have examined the genomic OR repertoire of 20 bird genomes spanning the avian phylogeny, revealing between 50 and 390 intact ORs per species (Driver and Balakrishnan, subset shown in Fig. 1). This suggests a more complex sense of smell in birds than previously appreciated (Driver and Balakrishnan 2021). However, we cannot discern the functional roles of ORs in smell without expression studies of the bird OE. I hypothesize that across the bird phylogeny, species express different subsets of their OR repertoire in the OE. Further, I hypothesize the some ORs are consistently present or absent from the OE, and that absent ORs are possible nonfunctional or expressed in other tissues.
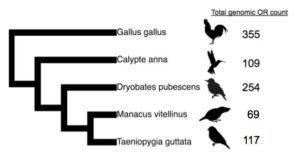
Using individual birds that were already sacrificed as part of ongoing projects, I have recently dissected OE and liver from the chicken (Gallus gallus) and zebra finch (Taeoniopygia guttata). I also intend to obtain OE samples from Anna’s hummingbird (Calypte anna), downy woodpecker (Dryobates pubescens), and golden-collared manakin (Manacus vitellinus). These species span avian diversity and represent diverse ecology (Fig. 1). Additionally, I have surveyed the genomic OR repertoires to find that all five species have over 60 total intact ORs (Fig. 1). I will compare OR expression in the OE to the liver and identify expressed ORs in both tissues. I will formally compare OE expression between the five bird species OE to measure species-specific differences in expression. The AGA EECG award is funding the critical OE expression work, which has never been performed in birds in a phylogenetic context. In addition to standalone results characterizing OR expression in the bird OE, implicating which ORs in the genomic repertoire are involved in olfaction will allow for subsequentwork to functionally test the unknown binding properties of bird ORs.
References



