**The AGA grants EECG Research Awards each year to graduate students and post-doctoral researchers who are at a critical point in their research, where additional funds would allow them to conclude their research project and prepare it for publication. EECG awardees also get the opportunity to hone their science communication and write posts over their grant tenure for the AGA Blog. In the first in the series, our EECG awardees write about their research and their interests as an ’embarkation’.**

About the Blog Author: Dr. Christine Ewers is an evolutionary biologist and postdoc at the Zoological Museum of the Kiel University in Germany. She uses aquatic invertebrates to reconstruct anthropogenically driven change and life history evolution. Follow Christine on twitter @ewers_c or via her website.
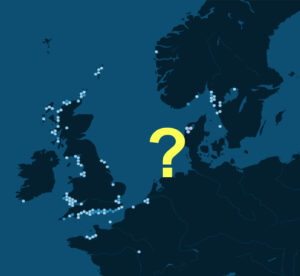
Oysters are not only an expensive delicacy, but also keystone species and ecosystem engineers. Unfortunately, they are threatened at a global scale by overfishing, pollution and climate change (Beck et al. 2011), which means their decline has tremendous impacts for foodies and coastal ecosystems alike. Like many other species, the European oyster Ostrea edulis L. has declined throughout its range in the past 150 years. Most drastically, it went locally extinct in the German and Danish Wadden Sea around 1930, presumably due to a combination of overfishing, harsh winters and disease. What I find curious is that it has not yet recolonized this habitat, even though fishing ceased decades ago, and much of the Wadden Sea is now a protected nature reserve (Fig. 1).
To understand the extinction and continued absence of European oysters from the Wadden Sea, we used museomics to sequence the genomes of 150-year-old oyster shells housed at the Zoological Museum of Kiel University (Fig. 2). Those data revealed that the Wadden Sea population had a distinct genetic makeup with one private mitochondrial haplogroup, and was significantly differentiated at the nuclear genome as well (Hayer et al. 2021, Ewers et al. in prep.). This indicates isolation of the Wadden Sea population, which could be caused by neutral processes, e.g. limited dispersal of the planktonic oyster larvae.

Whether dispersal is limited or not, the Wadden Sea population was also likely adapted to the challenging Wadden Sea conditions. In our soon-to-be-published work, we identified signatures of adaptation in the Wadden Sea population using genomic outlier approaches of Fst and Tajima’s D (Ewers et al. in prep.). If these adaptations went extinct with the Wadden Sea population, it would explain why the European oyster has not recolonised the Wadden Sea; the required functionality is no longer present in the species. However, if the Wadden Sea diversity survived somewhere, and has not recolonized the Wadden Sea due to limited dispersal, this remnant population could be used to experimentally investigate the causes of this local extinction event, and potentially restore the Wadden Sea ecosystem.
The Danish Limfjord represents a likely refuge of the Wadden Sea diversity. During a large storm in the 18th century, the Limfjord became connected to the North Sea through a narrow channel. In the following years, a large European oyster population became established (Möbius 1877), and represents one of the most pristine – and best-tasting – oysters in the world today. The exact origin of the Limfjord.

oysters is not known, but a colonization from the neighboring Danish Wadden Sea is most probable. A comparison of historical and present-day mitochondrial data supports this hypothesis: the Limfjord population is most similar to the historical Wadden Sea population. I will use the EECG grant to assess if the Danish Limfjord is a remnant of the extinct oyster diversity from the Wadden Sea.
References
Möbius, K. A. 1877. Die Auster und die Austernwirthschaft. – Parey.



