**The AGA grants EECG Research Awards each year to graduate students and post-doctoral researchers who are at a critical point in their research, where additional funds would allow them to conclude their research project and prepare it for publication. EECG awardees also get the opportunity to hone their science communication and write posts over their grant tenure for the AGA Blog. In the first in the series, our EECG awardees write about their research and their interests as an ’embarkation’.**

About the Blog Author: Peter Price is a PhD candidate with Dr Alison Wright at the University of Sheffield. Currently employing genomic techniques to explore the evolutionary dynamics acting on the transcriptome in the face of sexual selection. Keep updated with recent work on twitter @PeterDPrice
Sexual dimorphisms are one of the most pervasive forms of diversity across animals. Sculpted by sexual and natural selection, males and females can differ in a range of phenotypes, including form, colour, behaviour and more. These differences are driven both by differing reproductive roles of the sexes as well as inter- and intra-sexual competition, the latter of which can start an evolutionary cascade in which the exaggeration of traits, often a visual manifestation of the fitness of the male, leads to extreme dimorphisms. However, as most of the genome is shared across the sexes, the formation of sexual dimorphisms relies on some remarkable regulatory trickery of the genes present in both sexes. It is this dramatic display of alternative regulation that made me decide to join Alison Wright’s lab at the University of Sheffield to explore how a single genome can encode some of the most striking forms of diversity in the natural world.
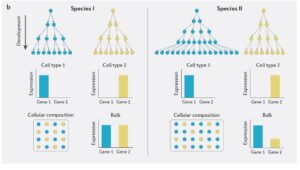
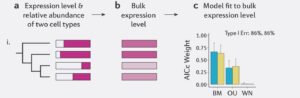
As an early PhD student, I started off exploring a variety of methods for testing how selection acts differently on expression in males and females, using birds as our model system. One challenge to current studies of expression evolution is that studies typically measure expression across entire heterogeneous tissues and compare this across species. This can make it difficult to distinguish changes in gene expression from differences in tissue composition (Figure1.jpeg: from Price et al 2022). Our recently published analysis (Price et al. 2022) showed, through some very simple simulations, that changes in tissue composition across species can have huge impacts on the inference of evolutionary processes. In extreme cases, we infer selection in the absence of any change in gene expression across species (Figure2.jpeg: from Price et al 2022).
One way to get around this challenge is by measuring gene expression patterns at the single-cell level, using single-cell RNA-seq (scRNA-seq). This state-of-the-art technology generates expression data at the single-cell resolution, and uncovers the cell types present in a tissue with their relative abundances, all with limited a priori knowledge. This is particularly useful to study the gonad, which is a target of strong sexual selection, and is therefore likely to exhibit large composition differences across species as a product of sperm competition (Lupold et al. 2009)
We are currently using scRNA-seq to explore how tissue composition varies both across development and across a range of avian species, by generating a comparative and temporal atlas of the avian gonads. In doing so, we are testing how sexual selection acts on gene expression and the role of regulatory change in sexual dimorphism. This AGA grant will support our ongoing work to establish the role of gene expression in adaptive change and phenotypic divergence. Hopefully, with this novel and exciting dataset, we’ll be able to test both how sexual selection acts on gene expression and the relative role of regulatory change in sexual dimorphisms.
References



