**The AGA grants EECG Research Awards each year to graduate students and post-doctoral researchers who are at a critical point in their research, where additional funds would allow them to conclude their research project and prepare it for publication. EECG awardees also get the opportunity to hone their science communication and write posts over their grant tenure for the AGA Blog. In the wrap-up to the series, awardees talk about their award and research in their ‘epilogue’.**

About the Blog Author: Dr. Claire Couch is a field ecologist and bioinformaticist and is currently a postdoc at Oregon State University. Claire studies commensal and pathogenic microbes in fish and wildlife systems and is interested in linking microbiome variation with host physiology, ecology, and population health.
I had the honor of receiving an EECG award relatively early in my PhD program to fund sequencing of nasal microbiome samples from a herd of African buffalo and assess drivers of microbiome variation, including relatedness and genetic variables. However, these nasal microbiome data were found to be extremely low-biomass and stochastic with respect to animal traits including relatedness and other genetic factors, therefore we do not anticipate that studies resulting from this dataset will be of interest to the Journal of Heredity readership. However, this grant funded one of my first independent research projects, and I learned a great deal from every part of the process, from the grant application to data collection and analysis. My experience with this study laid the groundwork for a number of other studies I have since successfully published on the subject of wildlife microbiomes. In the nearly five years since receiving the EECG award, I have contributed hypothesis-driven research of host-microbe interactions across scales using data from African buffalo (Syncerus caffer), desert bighorn sheep (Ovis canadensis nelsoni), Rocky Mountain elk (Cervus canadensis nelsoni), Chinook salmon (Oncorhynchus tshawytscha), and Pacific walrus (Odobenus rosmarus divergens).
Even though my original research project funded by the EECG did not go as planned, I wanted to make a meaningful contribution to the American Genetic Association in recognition of the important role that the grant played in my development as a scientist. One of the central questions that I study is how the microbiomes of wild animals interact with population genetic structure, environmental variation, and spatial structure in natural environments, and I believe this question may also be of interest to many in the conservation genetics community. As such, I took this opportunity to write a synthesis paper, with my population geneticist colleague Clint Epps, on applying population and landscape genetic approaches to gut microbiome research in wild populations (Couch & Epps 2022). In this work, we propose a conceptual framework for understanding wildlife gut microbiomes in relation to landscape variables and host population genetics, including the potential of approaches derived from landscape genetics. We use this framework to review current research, synthesize important trends, highlight implications for conservation, and recommend future directions for research. Specifically, we focus on how spatial structure and environmental variation interact with host population genetics and microbiome variation in natural populations, and what we can learn from how these patterns of covariation differ depending on host ecological and evolutionary traits.
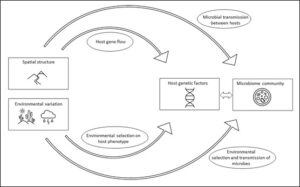
The gut microbiomes of natural host populations exhibit complex relationships with environmental, spatial, and host genetic factors (Figure 1). Because these variables are often highly correlated in natural populations, disentangling how they impact the microbiome is challenging, but is nonetheless key to understanding gut microbial ecology, addressing evolutionary questions, and identifying conservation and monitoring applications. This literature review and synthesis is framed around three questions with the goal of interpreting current knowledge and identifying gaps and future directions: 1. How do spatial structure and environmental variation mediate interactions between host population genetics and microbiome communities? 2. What are the relative effects of host genomic variation, spatial structure, and external environment on the microbiome? 3. How do the answers to (1) and (2) differ among host and environment categories?
Examining current research through the lens of these questions, we find that spatial, environmental, and host genetic structure can significantly impact gut microbiome composition and diversity, with potential consequences for wildlife health and conservation. Intriguingly, the relative influences of spatial, environmental, and host genomic variation appear to differ among host species and environments. However, the lack of wildlife microbiome studies that integrate host genetic data alongside spatial and environmental covariates currently limits our understanding of the underlying processes that shape microbiomes across spatially distributed host populations. Consequently, we lack theoretical bases for understanding and predicting how the microbiomes of wild populations may change in response to ecological, evolutionary, and anthropogenic change. To address these fundamental gaps, we recommend that researchers studying the microbiomes of wildlife populations consider incorporating information on host population genetic structure. These efforts may be assisted by analytical approaches from related fields such as landscape genetics. Ultimately, improved understanding of microbiome variation in natural populations could facilitate improved monitoring and management tactics for wildlife population health.
It is undoubtedly an exciting time to be involved in this new frontier of wildlife microbiome research, and I look forward to continuing to work with amazing colleagues across multiple disciplines to better understand the causes and consequences of microbial community variation in natural host populations. Again, I have the AGA to thank for helping to launch my career in this exciting field.
References



