About the Author
 Chris Robinson (he/him) is a PhD candidate in Bob Cox’s lab at the University of Virginia. His interests lay in how hormones contribute to phenotypic development and the evolution of hormone-genome interactions. His work uses hormonal manipulations, transcriptomics, and cellular imaging to understand how traits are gained and lost among closely related species. Chris loves to run and climb and believes that these hobbies are advantageous for being a capable lizard catcher!
Chris Robinson (he/him) is a PhD candidate in Bob Cox’s lab at the University of Virginia. His interests lay in how hormones contribute to phenotypic development and the evolution of hormone-genome interactions. His work uses hormonal manipulations, transcriptomics, and cellular imaging to understand how traits are gained and lost among closely related species. Chris loves to run and climb and believes that these hobbies are advantageous for being a capable lizard catcher!
Check out his new paper here!
Hormones contribute to the development of many different traits, often by activating hormone receptors that alter gene expression. When a hormone is circulated at different concentrations between females and males, sexually dimorphic traits can develop. Past research has sought to understand the evolution of sexually dimorphic traits, often by investigating how changes to circulating hormone levels regulate sex-specific trait development. However, the regulation of trait development by hormones is complex, and many intermediate steps can evolve to produce novel patterns of sexual dimorphism among closely related species (Cox et al. 2022). In this EECG-funded paper, we used a combination of hormone manipulation experiments and RNA sequencing in two species of lizards with different patterns of sexual coloration to test for species-level differences in tissue and gene sensitivity to hormonal stimulus.
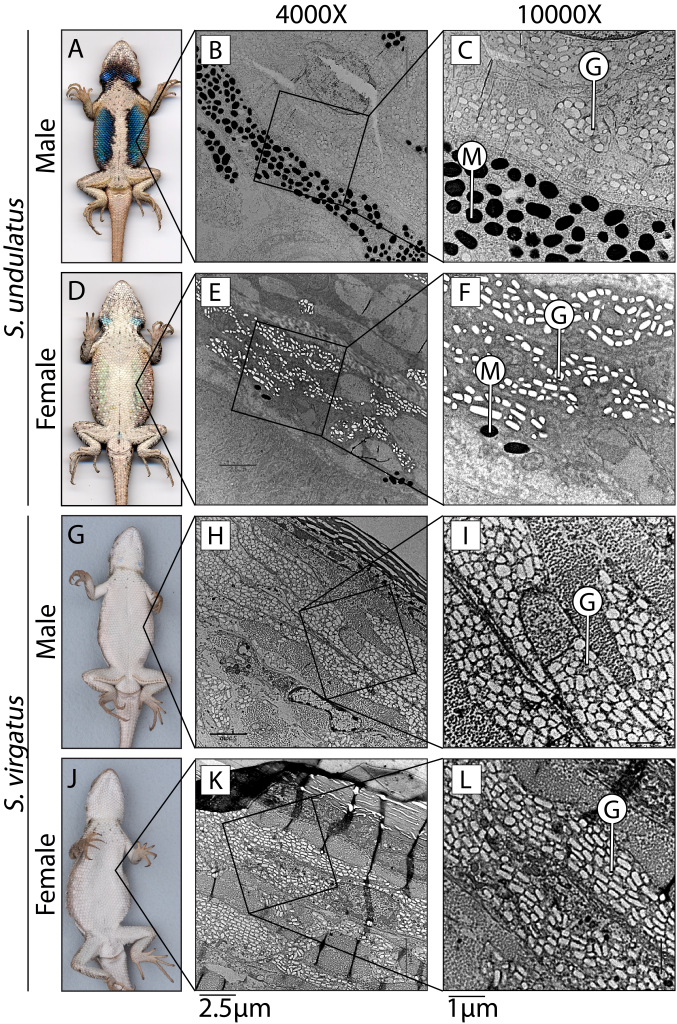
Lizards in the genus Sceloporus serve as an excellent model for addressing the evolution of hormone-phenotype couplings because 1) testosterone induces the development of sexually dimorphic phenotypes (e.g., Cox et al. 2005), and 2) the evolution of sexual dimorphism is remarkably labile within the genus. In terms of coloration, the ancestral state (Fig. 1A,D; S. undulatus) is represented by vibrant ventral coloration specific to males. However, males have independently lost this coloration approximately 13 times (Wiens 1999), resulting in a derived sexually monomorphic state (Fig. 1G,J; S. virgatus). A major difference between these species is the production of melanin in response to testosterone (e.g., Quinn and Hews 2003; compared Fig. 1C and Fig. 1I). Therefore, we had two foci for this project. First, we tested whether genes that regulate tissue sensitivity to a hormone differed between these species. Specifically, we examined whether ventral skin differed in the number of genes responding to testosterone between these species, and whether genes that affect androgen signaling (such as by converting testosterone into the more potent dihydrotestosterone, or expression of the androgen receptor itself) explained any observed differences. Second, we tested whether the expression of genes related to melanin synthesis differed in their response to testosterone to examine whether the sensitivity of specific genes regulated phenotypic differences (Fig. 2).
We found that ventral skin in the sexually dimorphic S. undulatus was substantially more responsive to testosterone than the sexually monomorphic S. virgatus, not because of differences in androgen receptor expression (surprisingly, the sexually monomorphic S. virgatus has higher androgen receptor transcript abundance!), but potentially through greater conversion of testosterone to dihydrotestosterone. We also found that the production of α-melanocyte stimulating hormone (α-MSH) and melanin synthesis was androgen-regulated in S. undulatus, but not in S. virgatus. Together, these results suggest that the evolution of sexual dimorphism in this system is associated with changes in gene sensitivity downstream of androgen signaling, with regulation of tissue sensitivity potentially mediating this relationship.
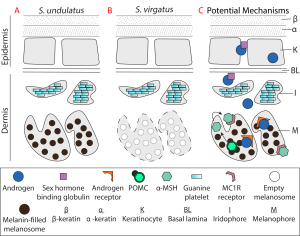
Our next step to understanding the evolution of sexual dimorphism is this system is to use immunohistochemistry to characterize protein abundance and localization in ventral skin of adults of these species. Cellular abundance can affect RNAseq data (Darolti and Mank 2023), so we want to examine whether the observed differences in the transcriptome can be attributed to differences in cellular architecture between the species. Our data suggest that melanophores or their precursors are equally abundant between S. undulatus and S. virgatus, as indicated by the expression of the genes MITF, DCT, and KIT, but the stage and/or activity of these cells cannot be inferred from our data. Further, if the cell type in which the androgen receptor is expressed differs, this could explain why melanin is synthesized by testosterone only in S. undulatus. This experiment offers an avenue to more deeply explore some of the complex relationships elucidated by our RNAseq data. Further, it adds to the emerging perspective in evolutionary endocrinology that hormone-phenotype couplings and hormone-genome interactions are an important avenue to examine when differences in hormone concentrations do not fully explain evolutionary phenomenon.
We would like to deeply thank the American Genetic Association for allowing us to explore this topic and look forward to sharing new results soon!
References
Cox RM, MD Hale, TN Wittman, CD Robinson, and CL Cox. 2022. Evolution of hormone-phenotype couplings and hormone-genome interactions. Hormones and Behavior 114:105216.
Robinson CD, MD Hale, TN Wittman, CL Cox, HB John-Alder, and RM Cox. 2023. Species differences in hormonally mediated gene expression underlie the evolutionary loss of sexually dimorphic coloration in Sceloporus lizards. Journal of Heredity 116:637-653.
Cox RM, SL Skelly, A Leo, and HB John-Alder. 2005. Testosterone regulates sexually dimorphic coloration in the eastern fence lizard, Sceloporus undulatus. Copeia 2005:597-608.
Wiens JJ. 1999. Phylogenetic evidence for multiple losses of a sexually selected character in phrynosomatid lizards. Proceedings of the Royal Society of London B: Biological Sciences 266:1529-1535.
Quinn VS and DK Hews. 2003. Positive relationship between abdominal coloration and dermal melanin density in phrynosomatid lizards. Copeia 2003:858-864.
Darolti I and JE Mank. 2023. Sex-biased gene expression at single-cell resolution: cause and consequence of sexual dimorphism. Evolution Letters 7:148-156.



