**The AGA grants EECG Research Awards each year to graduate students and post-doctoral researchers who are at a critical point in their research, where additional funds would allow them to conclude their research project and prepare it for publication. EECG awardees also get the opportunity to hone their science communication and write posts over their grant tenure for the AGA Blog. In the wrap-up to the series, awardees talk about their award and their research in their ‘epilogue’.**

About the Blog Author: Mike (he/him) is a PhD candidate at Florida State University working in the lab of Dr. Darin Rokyta. He is broadly interested in understanding how natural selection shapes predatory traits at the molecular level. While his background emphasizes field work and organismal research, his current focus is deciphering complex traits using genomics, transcriptomics, and other sequence-based bioinformatic strategies. He was awarded the EECG in 2021 for his work characterizing sensory phenotypes and investigating the coevolution of venom and sensory perception in rattlesnakes. Follow Mike on twitter @Mike_Hogan_
Snakes, like many smaller-bodied animals, are often considered secretive. While this idea represents the perspective of us scary humans (i.e., large, warm-blooded mammals), many aspects of their biology are simply invisible to us through behavioral observations alone. How do they “smell” their food or find a potential mate? How does venom toxicity change throughout their lifetimes? My PhD project attempts to fill some of these gaps using genomics, and receiving the AGA EECG award last year has directly supported me in realizing these goals.
Complex behaviors, such as predation, rely on multi-trait coordination at the level of the underlying genes. In rattlesnakes, predation requires sensory detection of prey followed by incapacitation by envenomation. After publishing my initial findings characterizing the complete chemosensory repertoire of the eastern diamondback rattlesnake (Crotalus adamanteus) last year (Hogan et al. 2021; free online view-only link here), I turned my attention towards annotating other genes, starting with venom. Interestingly, many of the rattlesnake venom genes occur in the genome grouped together in tandem repeat arrays, mirroring what I observed in the chemoreceptors. Conservative estimates suggest that ∼25% of all gene duplication events in vertebrates are tied to tandemly arrayed genes (Pan and Zhang 2008), suggesting gene duplications likely played a role in the diversification of rattlesnake venom and chemoreceptors. But how are these genes used? Can we detect regulatory overlap between venom and sensory systems? To begin understanding the molecular mechanisms coordinating venom and sensory phenotypes in the eastern diamondback, we needed to dig deeper than just the venom and chemosensory genes.
The utility of a genomic resource is inherently tied to the quality of the corresponding gene annotations. Most notably, predicting the functional significance of a gene product requires accurate recovery of gene protein-coding sequences. Up to this point, all our rattlesnake genes have been meticulously annotated by hand based on RNA-seq reads mapping out exonic structure. But, there may still be hundreds of additional non-venom, non-sensory genes called transcription factors that play crucial roles in regulating and maintaining these traits. To identify these targets, we chose to annotate all remaining genes in the genome, search for any that look like transcription factors, and finally test for regulatory activities linking them to our traits of interest. To give us the best possible chance of recovering accurate coding sequence annotations for the tens of thousands of remaining genes, we introduced machine learning bioinformatic tools into our pipeline. Specifically, we combined the best gene predictions using the MAKER (Holt and Yandell, 2011), BRAKER (Hoff et al., 2019), and AUGUSTUS (Hoff and Stanke, 2018) pipelines. Each of these methods relies on training an algorithm to “know what to look for” through reference-based machine learning. From our carefully selected high-quality training references, the neural network parameters driving these machine learning processes “learned” how to find and annotate the remaining genes in the genome.
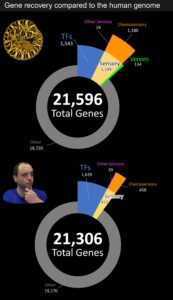
The final gene tally for the eastern diamondback genome came out to 21,596 genes, which is surprisingly close to what is predicted for the human genome (Figure 1). From the predicted coding sequences, we next implemented the DeepTFactor (Kim et al., 2020) pipeline to identify high-probability transcription factor coding genes based on previous deep machine learning. In total, we identified 1,543 candidate transcription factor genes, which again is surprisingly close to the number of transcription factors reported in humans (Figure 1). We plan to publish more on trait-specific details relating to these findings later this year.

Despite the COVID19 pandemic greatly limiting our fieldwork this past year, I was lucky enough to participate in a couple of collaborative field expeditions to sample venom and DNA samples from rattlesnakes on barrier reef islands. During these trips, I captured first-hand video footage of our fieldwork catching, measuring, sampling, and releasing wild rattlesnakes. With this footage, I put together a ~20-minute outreach video sharing our perspectives as rattlesnake biologists, which can be viewed on YouTube. I believe sharing these experiences directly from the scientists is crucial for making our research approachable to the public, especially for next generation youth scientists.
References
Pan D., Zhang L. (2008) Tandemly arrayed genes in vertebrate genomes. Comp Funct Genomics



