
About the Blog Author: Alexis Oetterer wrote this post as part of Dr. Stacy Krueger-Hadfield’s Scientific Communication Course at the University of Alabama at Birmingham. She is an PhD student in the Krueger-Hadfield evolutionary ecology lab and is interested in studying life cycle evolution and ecology. Follow Alexis on Twitter @AlexisOetterer.

The Salish Sea surrounding the San Juan Islands is home to an incredibly high diversity of seaweed species. However, climate change and ocean acidification mean that these lush algal habitats are changing. Some species once found in large numbers are now almost nonexistent. Some species are showing up – either from migrating to higher latitudes as sea surface temperature increases or being introduced through various ways. To track this change, we need a baseline of what species are present and taxonomic expertise in identifying them. Algae that are morphologically similar to native species may go unnoticed, as was the case with the marine invader Gracilaria vermiculophylla along the Eastern Shore of Virginia until the late 1990’s (Thomsen et al. 2006).

Within the first few days of arriving at Friday Harbor Laboratories for the Marine Botany: Diversity and Ecology summer course, I jumped into collecting seaweed. Armed with five-gallon buckets, bags, and rubber boots, Dr. Thomas Mumford led our class along a winding path to our first field site: Shore Trail (Figure 1).
As we explored the rocky intertidal zone, each student collected seaweeds that piqued their interest. After filling our buckets, we hauled them back to the lab where the flow-through seawater tanks were waiting.
Using a key to the seaweed species of the region, we processed and identified what we had collected based on morphological characteristics and cross sections. Dr. Mumford and Dr. Wilson Freshwater explained how to press seaweeds (Figure 2) and the importance of creating voucher specimens. Vouchers are preserved specimens that serve as a permanent record and are typically accessible in an herbarium collection.

Photo credit: D. Wilson Freshwater
Each student selected two or more macroalgae to barcode. Barcodes provide a species identification based on a short section of DNA from a specific gene. After DNA extraction and amplification, the samples were sequenced. We used the sequences to confirm species identity before uploading them to the Barcode of Life Data System.

Photo credit: D. Wilson Freshwater
In total, 39 pressed specimens were deposited at the University of North Carolina at Wilmington and the University of Washington herbarium, including my voucher specimens Ceramium californicum (Figure 3), which was a tiny red seaweed living epiphytically on a different alga, and Sarcodiotheca gaudichaudii (Figure 4). They are a record of what was found in the area at the time and can be used by future students and researchers. For instance, we were able to extract DNA from decades old pressings.
In addition to barcoding, we also worked in small groups to develop a research project. My project was focused on kelp. Kelps have a biphasic life cycle (Figure 5), with large sporophytes and microscopic gametophytes. Reproductive success in both phases of the life cycle is important since stages are linked (gametophytes produce egg or sperm and sporophytes produce spores that make new gametophytes), but little is known about settlement and recruitment of the gametophyte stage (Schoenrock et al. 2021). It is thought that coralline algae may play a role by providing settlement cues for the gametophytes.
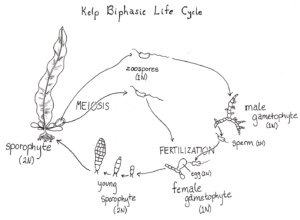
Illustration by Greta Bolinger
To test this, Greta Bolinger, Jordan Winter, and I collected samples of coralline algae found near kelp from Cattle Point. We identified three separate species of corallines: Bossiella plumosa, Calliarthron tuberculosum (Figure 6), and Corallina vancouveriensis. We examined the genicula, which are noncalcified, flexible regions on coralline algae (Figure 7), under a microscope and extracted DNA from samples containing potential gametophytes. Samples that amplified were sequenced, but each sample contained multiple DNA fragments, which made the kelp species potentially found in coralline algae unidentifiable. Using metabarcoding would allow for the identification of multiple taxa within a sample (Atika et al. 2020).

Photo credit: Greta Bolinger
My summer in the San Juans was one for the books…and for the field. After a year and a half of mostly online classes due to Covid, the Marine Botany course was a welcome opportunity to get back in the classroom, lab, and field. Everything I learned in the summer of 2021 has provided a solid foundation for my dissertation research.
Fast-forward a year later, and I am now a graduate student at the University of Alabama at Birmingham. In just a few short weeks I will be in the field again. Only this time, I will be on the East Coast, but still studying seaweeds and still trying to figure out what happens to gametophytes.
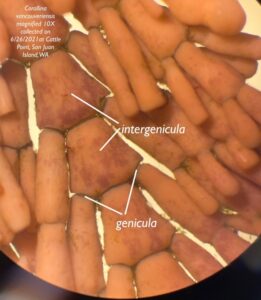
Photo credit: Greta Bolinger
References



