
About the Blog Author: Taylor Williams (she/her) is a Masters student in Dr. Heather Spalding’s lab at the College of Charleston. Her undergraduate degree is from the University of Hawai`i, Mānoa where she earned a B.S. in Marine Biology and first became a scientific diver. Since then, she has become an avid scientific diver and field scientist whose research has taken her to remote atolls of the Pacific. She uses population genetics to investigate the reproductive system of a cryptogenic seaweed that is acting invasively in a marine national monument.
The 15-foot, open ocean swell rocks and sways the ship, sloshing here and there. A dull ache in the back of your head signals looming sea sickness … and that’s just while you’re trying to sleep.
During the day, you’re never able to get quite dry enough. You’re cramped in the dive boat, a never-ending layer of salt on your skin, and a Rudolph-red nose because no matter how much sunscreen you apply, you’re just always one step behind the sun… and that’s just while you’re diving.
Once the dive day is over it’s time to rinse off your gear (wet again), take a shower (wet again), sort through the samples you collected (wet again), process all the samples (wet again), and then clean everything up (wet yet again), just in time to crawl into bed and sleep (ahhh … finally dry).
Why put ourselves through it? As they say in Hollywood: location, location, location.
The driving force for many of us is equal parts passion for the science and immense gratitude for traveling to far flung places to collect the data.
Papahānaumokuākea Marine National Monument (PMNM) encompasses the Northwestern Hawaiian Islands – the most remote island chain on the planet composed of a series of uninhabited atolls and seamounts (Figure 1). The remote location of this archipelago provides unique opportunities to address scientific questions that are often challenging to investigate.

Of course, there are also some perks tied to working in remote locations like PMNM – the sunrise, sunsets, and starry nights. The teal blue water filled with megafauna can’t be rivaled by any artist and being one of the handful divers to get a first-hand look at the many endemic species of the region is just the icing.
I’ve personally had the grueling pleasure of working within the boundaries of PMNM three times. For this past 2021 field season, we were particularly interested in addressing some population-level eco-evolutionary questions on a cryptogenic seaweed. Comparable population-level analyses within PMNM have primarily focused on fishes to assess connectivity within and outside the monument boundaries, particularly for management purposes (Rivera et al. 2011, Toonen et al. 2011).

My focus during both field seasons was the red seaweed Chondria tumulosa (Sherwood et al. 2020) (Figure 2) – a newly described species overgrowing the reefs around Manawai, or Pearl and Hermes Atoll (Figure 3). This species was first recorded in 2015 in small distinct patches. By 2019, thick mats had overgrown kilometers of the reef (Figure 4).
It’s recent establishment, high biomass, and remote island location within a marine national monument make this seaweed a target management species to try to understand and prevent its spread.
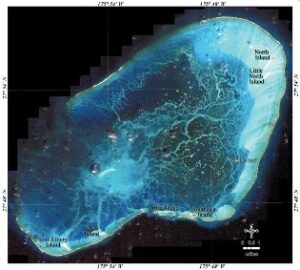
Studying C. tumulosa has proven challenging at nearly every step. Access to its only known location requires, at minimum, three weeks of ship time and an entire crew. It’s a hot topic threat to PMNM, so there’s enhanced biosecurity. Manawai is a four-day ship transit to the nearest inhabited island making sample preservation difficult. Once back in the lab, its haploid-diploid life cycle makes subsequent genetic work challenging because the methods used in diploid taxa aren’t quite as cut and dry for C. tumulosa.
Like other red seaweeds, C. tumulosa has long-lived gametophytes (haploid) and sporophytes (diploid), effectively separating meiosis and fertilization both spatially and temporally (Figure 5). This separation leads to unique eco-evolutionary consequences (Krueger-Hadfield 2020). For example, selfing can occur in haploid-diploid taxa even when there are separate sexes because cross-fertilization between male and female gametophytes that share a diploid parent is analogous to selfing in a hermaphroditic angiosperm or animal (Klekowski 1969). Additionally, asexual reproduction will result in the dominance of one stage and loss of the others. For example, in the haploid-diploid red alga Gracilaria vermiculophylla, diploid dominance has been observed throughout the invasive populations of the Northern Hemisphere (Krueger-Hadfield et al. 2016). Because meiosis occurs in the sporophyte stage, the gametophyte stage can be recovered under the appropriate conditions, completing the life cycle (Krueger-Hadfield 2020). However, if the population is only female or only male, it will be difficult to recover the life cycle (i.e., recover sexual reproduction).

Reproductive system variability directly affects invasion success and long-term establishment of a species (e.g., Baker 1955), but studies of haploid-diploid taxa that assess both ploidy stages are few and far between, especially for invasions (Krueger-Hadfield 2020). Incorporating this information is essential for understanding the population dynamics of haploid-diploid taxa. The reproductive system also strongly influences both the genetic diversity and evolutionary ecology of a population (Barrett 2011). This is particularly true for haploid-diploid seaweeds for which this information has been largely overlooked due to the care that has to go into sampling (Krueger Hadfield & Hoban 2016) and the complexity of the lab work (Stoeckel et al. 2021).

Aside from knowing that C. tumulosa is a haploid-diploid species and some early preliminary visual assessments for reproductive structures, we know close to nothing about the reproductive system of this new invasive-like seaweed. Having this basic information will be an asset to all future studies on this species because it will provide some evolutionary context to future work. Luckily this field season provided us with a wealth of new samples that will allow us to investigate this species further and open the door to our understanding of its eco-evolutionary potential.
The 2021 field season is over. We successfully collected upwards of 800 samples! So, now it’s time to hit the lab and make up for the lost time of 2020. However, population-level work in haploid-diploid species isn’t always straight forward because the tools and techniques that have been developed for diploid-dominant species cannot be readily transferred to haploid-diploid taxa (Krueger Hadfield & Hoban 2016). Luckily, Stoeckel et al. (2021) and Krueger-Hadfield et al. (2021) recently provided a concise framework for addressing these haploid-diploid predicaments. The structure of haploid-diploid populations is highly influenced by the haploid proportion and rate of clonality (Stoeckel et al. 2021). If the population lacks gametophyte individuals completely due to fragmentation, analyses can still be conducted on the sporophyte thalli alone. Fragmentation is the hypothesized reproductive mode for C. tumulosa as it has been found to occur heavily at all sites. If this hypothesis is correct C. tumulosa will be another taxon in which we can explore the population genetic consequences of losing a free-living stage entirely.
I guess all that’s left now is to say farewell to my sea legs, trade in my dive gear for some pipettes, and get back to work.
Acknowledgements: A huge thank you goes out to the National Fish and Wildlife Foundation for providing the funding to make this work possible and to NOAA’s Papahānaumokuākea Marine National Monument team for continuously providing us access to this treasured slice of the planet.
References
Rivera MAJ, Andrews KR, Kobayashi DR, Wren JLK, Kelley C, Roderick GK & Toonen RJ (2011) Genetic Analyses and Simulations of Larval Dispersal Reveal Distinct Populations and Directional Connectivity across the Range of the Hawaiian Grouper ( Epinephelus quernus ). Journal of Marine Biology 2011:1–11.



