The ability of organisms to colonize or adapt to new and changing environments is of critical interest in biology, especially considering the threats and advances of human-induced environmental change. Certain traits may play a key role in determining population persistence or decline in the face of climate change, habitat fragmentation, and the like. In the Boughman Lab at Michigan State University, we study the role of sensory systems in habitat colonization, as they are important for many social and ecological interactions such as reproduction and foraging. We study the adaptation and diversification of sensory morphology (among other measures), including olfaction, vision, and mechanosensation in the Icelandic threespine stickleback (Gasterosteus aculeatus).
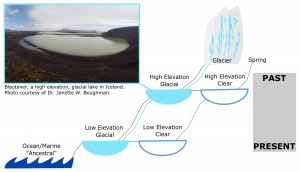
Threespine stickleback are found in high and low elevation lakes that differ in age across the dynamic landscape of Iceland, and these lakes also differ in their sensory ecology. Some lakes found at both high and low elevation are fed by glaciers and are blue-shifted and highly turbid, while others are spring-fed and are clear. Therefore, the lakes of Iceland and their stickleback offer a unique system in which to study sensory systems at different evolutionary time points and under different sets of ecological challenges. In order to better understand the role of sensory systems in adaptation and colonization, we also need to know the evolutionary history of, or the extent to which genetic variation and phenotypic plasticity contribute to sensory systems in threespine stickleback.
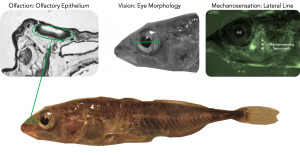

The Boughman lab is working to address the genetic variation underlying sensory systems and the genetic relationships between 25 different populations of threespine stickleback using a reduced-representation genome sequencing method called double digest RAD-tag sequencing (ddRADseq; Peterson et al. 2012). This technique uses two restriction enzymes to cut up DNA into smaller fragments, which are then size-selected for sequencing to find single nucleotide polymorphisms (SNPs) used for genotyping. By knowing how populations from lakes at different elevations and with different ecologies are genetically related, we can understand the rates at which any difference in sensory systems we measure have evolved. Threespine stickleback in Icelandic lakes all originated from the sea, so we will be able to construct their evolutionary history by comparing lake populations to extant “ancestral” marine populations. In general, we expect lakes at higher elevation to have been colonized earlier than lakes at lower elevation. We may expect stickleback from glacial lakes at different elevations to share similar sensory morphology, perhaps investing more in olfaction and mechanosensation over vision, but the amount of phenotypic change to be proportional to population age.
Phenotypic plasticity may contribute to and interact with natural selection to facilitate sensory system adaptation to novel environments. Epigenetic modifications (DNA methylation, histone modifications, noncoding RNAs) and epigenetic inheritance may underlie plastic responses, provide selectable variation and dictate evolutionary trajectories. DNA methylation, in particular, is a dynamic, heritable epigenetic modification in vertebrates that influences gene expression and may be critical to mediating plasticity and its facilitation of adaptation. We might expect that the methylated regions across the genome will vary with relative population age. For instance, we may expect our marine ancestral population to exhibit more or less methylation as compared with more derived, glacial freshwater populations. If “ancestral” marine populations have a greater potential for phenotypic plasticity than glacial, freshwater populations, we may expect fewer methylated regions across the genome, as methylation typically represses gene expression. With funding from an AGA Evolutionary, Ecological, or Conservations Genomics Award, I will characterize epigenetic relationships among the same populations of stickleback used in the ddRADseq experiment using a technique developed by Schield et al. (2016) called EpiRADseq.
EpiRADseq is similar to ddRADseq except that one of the two restrictions enzymes is methylation sensitive, whereby fragments of DNA are produced and sequenced only when cytosine nucleotides at recognition sites are not methylated. Using this method, Schield et al. (2016) demonstrated dramatic transgenerational effects of predation on DNA methylation in water fleas (Daphnia ambigua). Recently, Crotti et al. (2020) tested whether EpiRADseq can, like ddRADseq, generate SNPs for population genomics in European whitefish (Coregonus lavaretus), and found that SNPs generated using each method were similar (55% of SNPs were shared between the two methods).
By using both methods, ddRADseq and EpiRADseq, we can first validate differences in methylation found with EpiRADseq; that is, we can know whether SNPs generated by EpiRADseq are indeed due to differences in methylation states rather than differences in genotype. Further, with our 25 populations of threespine stickleback, we can use SNPs to describe both the genetic and epigenetic relationships among populations that may vary with relative population age (lake elevation) and with ecology (lake environment).
Depending on the genetic or epigenetic sources of variation underlying differences in sensory systems, we may find that each method relates populations to each other differently. For instance, stickleback from lakes that are similar in age with but with different environments might be more divergent epigenetically than they are genetically. Differences in DNA methylation among populations may also indicate differences in the role of phenotypic plasticity in shaping sensory systems, with more methylated sites indicating less plasticity. The role of epigenetic modifications in shaping diversification and evolutionary trajectories is an exciting research frontier. This work will broaden our understanding of the interplay between epigenetics and adaptive evolution.
References:

Whitley Lehto is a postdoctoral research associate in Dr. Janette Boughman’s lab at Michigan State University where they are characterizing the genetic and epigenetic relationships among populations of threespine stickleback in Iceland from lakes that differ in their sensory challenges. Whitley completed their PhD at the University of Denver in 2019. They are interested in the ways in which phenotypic plasticity and epigenetic modifications may contribute to rapid population responses in new and changing environments.




