**This post is a part of the series on the 2019 AGA Presidential Symposium – Sex and Asex: the genetics of complex life cycles**
About the Authors:
 Sarah Shainker (she/her/hers) is a PhD student in Dr. Stacy Krueger-Hadfield’s lab at the University of Alabama at Birmingham. She uses population genetics to research the mating systems and eco-evolutionary dynamics of freshwater red algae in the eastern United States. She completed a B.S. in Marine Biology at the College of Charleston, where she became interested in evolutionary ecology, before serving with Peace Corps Philippines where she gained experience and interest in science communication, education, and outreach. Sarah tweets at @SarahShainker.
Sarah Shainker (she/her/hers) is a PhD student in Dr. Stacy Krueger-Hadfield’s lab at the University of Alabama at Birmingham. She uses population genetics to research the mating systems and eco-evolutionary dynamics of freshwater red algae in the eastern United States. She completed a B.S. in Marine Biology at the College of Charleston, where she became interested in evolutionary ecology, before serving with Peace Corps Philippines where she gained experience and interest in science communication, education, and outreach. Sarah tweets at @SarahShainker.
 Sabrina Heiser (she/her/hers) is a PhD Candidate in Dr. Charles D. Amsler’s lab at the University of Alabama at Birmingham. Her research focuses on the factors driving the geographic distribution of chemical defenses in a red seaweed. One of her projects is looking at the population genetics of that red seaweed which exhibits a haplodiplontic life cycle as discussed in the blogpost. For her sample and data collection, she gets to go and SCUBA dive in Antarctica. She received her B.Sc. in Marine Biology from Plymouth University (UK) and is originally from Germany. Follow her adventures on her website or @SabrinaHeiser.
Sabrina Heiser (she/her/hers) is a PhD Candidate in Dr. Charles D. Amsler’s lab at the University of Alabama at Birmingham. Her research focuses on the factors driving the geographic distribution of chemical defenses in a red seaweed. One of her projects is looking at the population genetics of that red seaweed which exhibits a haplodiplontic life cycle as discussed in the blogpost. For her sample and data collection, she gets to go and SCUBA dive in Antarctica. She received her B.Sc. in Marine Biology from Plymouth University (UK) and is originally from Germany. Follow her adventures on her website or @SabrinaHeiser.
While studying red macroalgae, we have learned that whether they inhabit Alabama’s streams or Antarctica’s subtidal forests, these organisms are more complicated than they may appear at first glance. Along with many other taxa, including mosses, ferns, algae, and some fungi, they have complex haplodiplontic life cycles that are wholly different to that of animals or the alternations of generations found in angiosperms. In addition to a unique life cycle, these organisms often have diverse reproductive strategies. These characteristics present an opportunity, as they provide an alternative lens through which to view predictions about the evolutionary maintenance of sex, which have mostly been made in more well-known organisms (i.e., angiosperms). However, this opportunity comes with a challenge: which population genetic tools can be used to accurately assess reproductive systems, because those commonly used for diplontic organisms cannot be directly translated to haplodiplontic ones.
Life cycles, together with their reproductive mode (sex vs. clonality, including partial clonality) and mating system (outcrossing vs. inbreeding, including mixed mating) play a major role in partitioning the genetic diversity of populations. In sexual populations and where unrelated individuals mate, we expect to find more diversity. In contrast, we tend to find lower genetic diversity in asexual populations or those populations in which individuals are inbreeding. These rules may seem relatively simple, however, there are some exceptions that make things a little more complicated.
Most eukaryotes, are not limited to sex. Instead, they often alternate between sexual and asexual reproduction (= clonality). Partial clonality has profound impacts on species’ ecological success and their ability to evolve in response to environmental change (Orive et al. 2017).
Most population genetic studies of partial clonality have focused on diplontic or diploid-dominant taxa (e.g., animals or flowering plants, respectively). Reduced genotypic diversity, higher rates of linkage disequilibrium, and a higher variance in Fis, including an excess of heterozygotes, are typically used as population genetic signatures of asexual reproduction (Stoeckel et al. 2021).
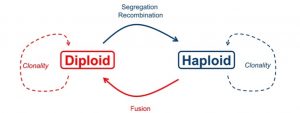
In haplodiplontic life cycles, both haploid and diploid stages undergo somatic development, resulting in both the spatial and temporal separation of meiosis and fertilization (Figure 1). This separation has profound evolutionary implications. For example, under full clonality, the haploid and diploid pools within a haplodiplontic population will become genetically differentiated, with the potential loss of one of the ploidy stages (Stoeckel et al. 2021). Additionally, a prolonged haploid stage provides a window during which both genetic drift and the purging of deleterious mutations can occur (Mable & Otto 1998). This all means that the proportion of haploids in a haplodiplontic taxon has implications for the genetic makeup and evolutionary trajectory of that population (Stoeckel et al. 2021).
Population genetic indices that are informative in organisms with diplontic life cycles may not be as useful in organisms with haplodiplontic life cycles. Moreover, not all population genetic indices used for diploid organisms can be used for haploids, and some indices will be skewed for diploids in haplodiplontic poulations. For example, heterozygosity cannot exist in haploid organisms because there is only one allele per locus instead of two. Additionally, although Fis is typically used as a measure of clonality in diplontic populations, it is less reliable in haplodiplontic populations because of noise caused by haploids (Stoeckel et al. 2021).
Stoeckel et al. (2021) simulated haplodiplontic populations for 10,000 generations. The results are influenced by the haploid proportion and by the clonal rate, so, what to do?
First, indices should be measured in the entire population of haploids and diploids as well as individually in each ploidy pool. Previous studies have typically assessed either haploids or diploids alone. Second, Fis,the metric most commonly used as a proxy for clonality in diploid-dominant populations, is not equally informative in haplodiplontic organisms because it is highly influenced by the proportion of haploids present in a population. However, pareto β may be an alternative metric that has rarely been calculated for haplodiplontic taxa, but is highly informative of clonal rates (Stoeckel et al. 2021).
The results of the simulations also indicated that partial clonality acts as a homogenizing force between haploid and diploid subpopulations, by maintaining genetic diversity from past generations while allowing for admixture between haploids and diploids. This finding suggests that partial clonality may be key to the evolutionary maintenance of haplodiplontic life cycles (Stoeckel et al. 2021).
Nature is much more complicated than any simulation. So what about the real deal – how do these predictioncs compare to real live, partially clonal haplodiplontic populations?
Studies investigating the reproductive system variation by genotyping and assessing patterns in both ploidy stages are rare in haplodiplontic taxa. Krueger-Hadfield et al. (2021) surveyed the literature for population genetic studies on mosses and seaweeds (none were available for ferns where both haploids and diploids were genotyped) and recalculated population genetic indices following recommendations made by Stoeckel et al. (2021). No studies had previously calculated pareto β, but Krueger-Hadfield et al. (2021) used this metric to categorize the clonal rates of species (Figure 2), and even populations within a species. Figure 2 shows the 14 species investigated from highly clonal (Phymatolithon calcareum) to highly sexual (Chondrus crispus).
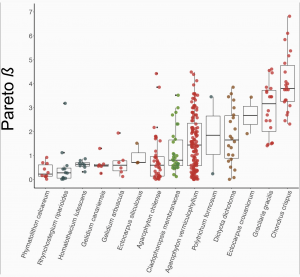
As predicted by Stoeckel et al. (2021), the size of haploid subpopulations contributes to genotypic richness because having more clones results in a lower number of genotypes within the population (Krueger-Hadfield et al. 2021). The prediction that genetic differentiation between life stages in highly clonal and highly sexual populations is higher than that of partially clonal populations was also confirmed (Krueger-Hadfield et al. 2021).
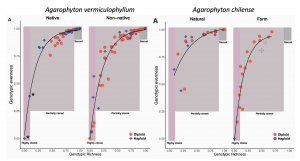
The authors delve into a finer scale exploration of two species of red seaweeds – Agarophyton vermiculophyllum and A. chilense. Agarophyton vermiculophyllumhas successfully invaded soft bottom habitats throughout the Northern Hemisphere and A. chilense has been farmed in similar soft-sediment habitats in Chile. In both cases, these free-floating populations are dominated by diploids through clonal, vegetative fragmentation (Guillemin et al. 2008, Krueger-Hadfield et al. 2016). In A. vermiculophyllum, populations in the non-native range exhibit more variation in clonality and a higher frequency of partial clonality (Figure 3). The same is true for A. chilense when comparing naturally occurring and farmed populations (Figure 3). However, A. chilense has no farmed populations considered sexual.
In order to accurately assess the population genetic structure and life cycle dynamics of haplodiplontic taxa, we have to account for all members (haploids and diploids) and use the appropriate indices. Krueger-Hadfield et al. (2021) called for calculating not only pareto β, but also the probability of identity between sibs and both the single-locus values and variance of Fis.
We clearly need more studies to further test the recommendations made by Stoeckel et al. (2021) and supported by Krueger-Hadfield et al. (2021). The context of these future studies will give us insight into how different environments interact with haplodiplontic life cycles and the eco-evolutionary consequences. Most studies appear restricted to terrestrial or temperate marine environments. Can the same patterns be found at higher latitudes (e.g., Antarctic marine algal forests)? Do freshwater red algae that inhabit flowing water, with ploidy stages that are physically connected to one another like mosses, display similar patterns? We hope to use the recommendations of Stoeckel et al. (2021) and Krueger-Hadfield et al. (2021) to find out!
References



