**The AGA grants EECG Research Awards each year to graduate students and post-doctoral researchers who are at a critical point in their research, where additional funds would allow them to conclude their research project and prepare it for publication. EECG awardees also get the opportunity to hone their science communication and write posts over their grant tenure for the AGA Blog. In another post in the series, awardees talk about their award and research in their ‘extension’.**

About the blog author: Anna O’Brien is joining the faculty Department of Molecular, Cellular, and Biomedical Sciences at the University of New Hampshire in March 2022, but is currently a Post-doctoral fellow at the University of Toronto in Ecology and Evolutionary Biology (EEB). Anna works in the labs of Profs Megan Frederickson (EEB), Chelsea Rochman (EEB), and David Sinton (Mechanical Engineering), and collaborates with the lab of Prof. Stephen Wright (EEB). Anna’s interests encompass species interactions broadly, but she primarily works on plant-microbe associations. Abiotic environments can shift costs and benefits of these interactions for both parties, and Anna’s post-doctoral work has shown that novel anthropogenic contaminants may often disrupt reciprocal benefits of plant-microbiome interactions. Follow Anna on twitter.
Organisms, especially aquatic species, often have physiologies finely tuned to salinity levels in their habitat (Brennan et al., 2018). Yet, the exposure of freshwater life to salty water is becoming commonplace around urban and suburban temperate zones (Dugan et al., 2017), where the application of salt as a de-icer to make roads and sidewalks safer also delivers salt to streams and groundwater in thaw events. In Toronto, the use of salt for de-icing has added so much salt to local ecosystems that streams are salty year-round (Lawson & Jackson, 2021).

Duckweed (Lemna minor) is a tiny, floating freshwater plant that primarily reproduces clonally via budding of tiny plantlets. In Ontario, duckweed thrives in still waters in pristine wilderness, but also thrives in urban Toronto where it is regularly exposed to salty runoff. How does duckweed manage this high range of salt tolerance?
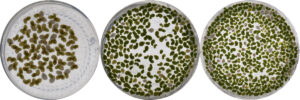
Figure 2: Experimenal duckweed in high salinity (5 g/L NaCl), showing less salt tolerant and more salt tolerant duckweed from left to right. June, 2021. Image credit: Anna O’Brien
Other species have achieved locally adapted salt tolerance via genetic changes (Brennan et al., 2018). However, duckweed has very little genetic variation in Ontario, likely in part due to its highly clonal reproductive habit (Ho et al., 2017), and duckweed appear to acquire salt tolerance in only a few frond generations (O’Brien et al., 2019), suggesting that salt tolerance is instead a plastic trait.
A major mechanism of plastic stress tolerance in plants is induced epigenetic marks in genes and nearby regions, such as the methylation of DNA cytosines. Stress-induced differential methylation can alter gene expression and improve tolerance to stressors, including salt (e.g. Boyko et al., 2010). Most plants reset this type of epigenetic mark in between generations, but clonal plants often do not (Wilschut et al., 2016). This has led some to hypothesize that transmitted cytosine methylation may be a source of adaptive, heritable variation in clonal plants like duckweed that have wide environmental tolerances (Wilschut et al., 2016, Harkess et al., 2020).
We took a clonal line that had not been exposed to salt in years, and placed individuals in jars without salt, or with high levels of salt for 5 weeks. With our #EECG2020 Award, we then extracted DNA and obtained enzymatic methylation sequence from the individuals in both treatments. While we have only just now cracked this dataset open, we have already learned some interesting things about methylation in Lemna minor.
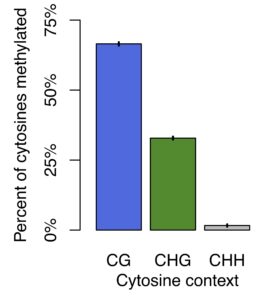
Cytosines in a strand of DNA will have different immediate bases as neighbors. This local context for a cytosine, such as whether the base immediately downstream is a guanine, changes which processes can methylate that cytosine. Methylation processes are thought to be highly conserved across plants. Yet, Harkess et al. (2020) recently discovered that a close relative of our duckweed, Spirodela polyrhiza, appears to have partial and complete loss of genes underlying some methylation processes, and as a result has very low methylation levels in particular contexts. In contrast, in our Lemna minor dataset, rates of methylation appear to be much higher than in S. polyrhiza. Preliminary analysis suggests an average of 66% of cytosines in CG context are methylated in L. minor, while in S. polyrhiza this number is only 10%. Rates of methylation in L. minor fall within the ranges observed for other plants, especially other monocots (Niederhuth et al., 2016). This implies unique losses of methylation in S. polyrhiza. Going forward, L. minor and S. polyrhiza, have similar ecology and often co-occur in the same ponds, providing a unique opportunity to study both mechanisms and eco-evolutionary consequences of a major shift in genomic regulation.
References



