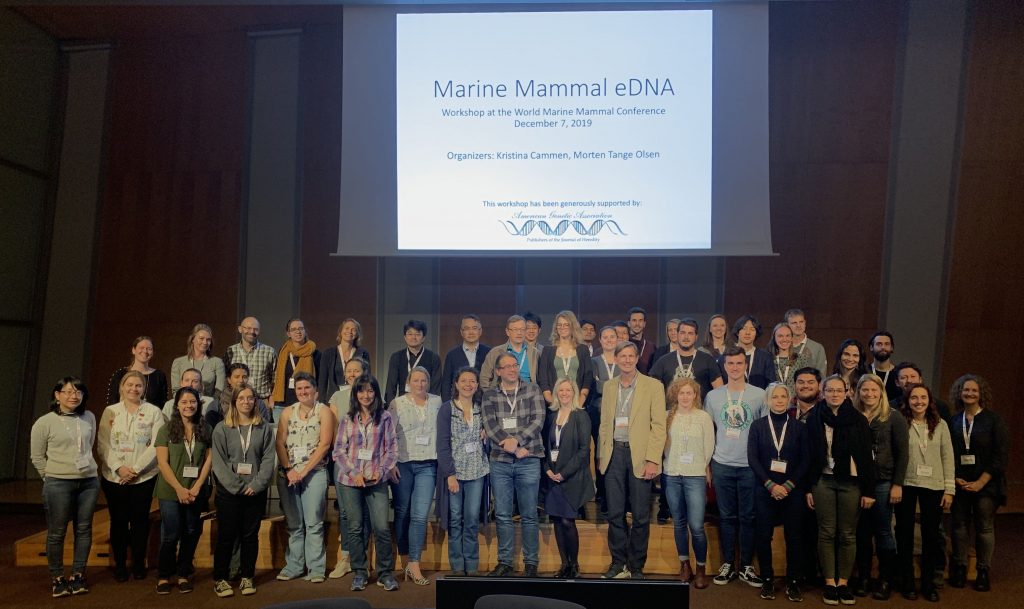
With a new set of tools focused on environmental DNA, or eDNA, we can learn a lot! eDNA approaches are built on the premise that organisms leave a trace of DNA in the water they inhabit, for example, through sloughed cells, urine, and feces. We can, therefore, extract organismal DNA from a water sample without ever handling, or even seeing, the organism.
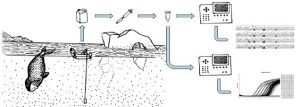
In brief, these approaches involve collecting a liter of water, filtering that water, and then extracting DNA from the filter. The risk of contamination in these highly dilute samples is high, so careful attention has been taken to develop appropriate protocols. These types of approaches are particularly appealing to marine mammal scientists, who hope to use eDNA to sample populations in a non-invasive, cost efficient manner.
With the support of an American Genetic Association Special Event Award, Kristina Cammen from the University of Maine and Morten Tange Olsen from the University of Copenhagen convened a one-day workshop on the topic of marine mammal eDNA at the recent World Marine Mammal Conference held in Barcelona, Spain in December 2019. The workshop included talks by six invited speakers and was attended by 60 participants. AGA funds were used to support the participation of early career scientists, including students and postdocs.

The workshop’s keynote speaker, Owen Wangensteen, kicked off the workshop by sharing his experience using metabarcoding techniques to assess eukaryotic biodiversity. Metabarcoding approaches use a single set of primers that are carefully selected to target a region of the genome that is shared across diverse species of interest and whose sequence is uniquely characteristic of species origin. Following amplification, researchers can use high-throughput sequencing to infer the origin of eDNA fragments from many different species present in a single water sample.
Alternative eDNA approaches focus on detection of a single species, rather than characterizing whole communities. Using highly sensitive quantitative approaches, such as qPCR and ddPCR, scientists can detect the presence of very low concentrations of free-floating DNA in water samples, for example water that is collected in the fluke print of a diving whale. Invited speakers Scott Baker, Maggie Hunter, Kim Parsons, Dóra Székely, and Kathryn Stewart provided examples of successful application of eDNA approaches to detect killer whales (Baker et al. 2018), beaked whales, manatees (Hunter et al. 2018), harbor porpoise (Parsons et al. 2018), bowhead whales, and the Yangtze finless porpoise (Ma et al. 2016).

Inspired by the science that the speakers presented and energized by a Catalonian lunch and coffee, workshop participants engaged in breakout group discussions for the remainder of the workshop day. Groups discussed aspects of eDNA science such as how to design an eDNA study, sample eDNA in the field, and set up an eDNA lab. Participants considered uses of eDNA for detecting species presence, as well as characterizing intra- and inter-specific genetic diversity of sampled populations and communities. Finally, a group tackled the challenge of how to analyze eDNA data, which involves both the adaptation of existing analysis pipelines, as well as the development of new statistical and analytical approaches.
At the end of the day, the feeling among workshop participants was one of cautionary excitement. These new approaches that leverage non-invasive sampling of eDNA hold a lot of promise for our field, but continued validation studies are needed. We are excited to see where this field goes next and anticipate great science coming from both early-career and more established scientists who participated in this workshop.
References

Kristina Cammen is an Assistant Professor of Marine Mammal Science in the School of Marine Sciencesat the University of Maine. In her research, Kristina uses primarily molecular approaches to study marine mammal population ecology and health. Over the past year, she has begun developing approaches for non-invasive sampling of genetic material from marine mammals using eDNA, with a particular interest in sampling water near seal haul-out sites. You can learn more about the work of Kristina and her students on her website.




