**This post is a part of the series on the 2019 AGA Presidential Symposium – Sex and Asex: the genetics of complex life cycles**
 About the author: Kamil S. Jaron is a Postdoctoral research fellow at the University of Edinburgh. Kamil is interested in genome evolution of species with unusual reproduction. His previous work with Tanja Schwander focused on large scale genomic consequences of parthenogenesis. He is currently working with Laura Ross on the Paternal Genome Elimination. As a by-product of his genomic analyses he co-developed smudgeplot, a visualization technique for understanding genome structure and ploidy. More to be found on his webpage or twitter (@KamilSJaron).
About the author: Kamil S. Jaron is a Postdoctoral research fellow at the University of Edinburgh. Kamil is interested in genome evolution of species with unusual reproduction. His previous work with Tanja Schwander focused on large scale genomic consequences of parthenogenesis. He is currently working with Laura Ross on the Paternal Genome Elimination. As a by-product of his genomic analyses he co-developed smudgeplot, a visualization technique for understanding genome structure and ploidy. More to be found on his webpage or twitter (@KamilSJaron).
We live in a wonderful era for evolutionary biology. With the advent of affordable sequencing we are getting hands on so many and so amazingly diverse genomes.
I am interested in the interplay of genome evolution with reproduction modes, so watching the growing body of literature with parthenogenetic genomes made me very happy. However, I noticed two striking aspects. First, many asexual genomes were unusual. Second, authors very often used one or two asexual species to propose some general effects of parthenogenesis. For example, in two parthenogenetic genomes (the springtail F. candida and the bdelloid rotifer A. vaga) genomic palindromes were reported. However not many other studies of asexual genomes looked into this genomic feature. Are palindromes widespread? Or has the absence of palindromes been underreported? Similar to the palindrome example, many genomic features, including features of key interest for the evolution of parthenogenetic species such as heterozygosity, were not systematically studied in the individual genomes.
We started to look at some of the asexual genomes on our own, before we realized that it would be worth analyzing them all, which later became our contribution to this special issue – Genomic Features of Parthenogenic Animals. We collected both reported genomic features (Figure 1 in the paper) but also the sequencing data and estimated as many relevant features as possible. When we started collecting, about 15 parthenogenic genomes were available, but soon asexual genomes started popping up at an incredible pace of a genome per month or two. We had to design a way to analyze all the new genomes automatically. Our pipeline was designed for automated downloading of sequence data and recalculating most of the analyses on the fly, as it would otherwise be impossible to keep up with all the new data coming in (and the workflow is openly available).
We were really proud to pull this off, and with all the glory, we posted a preprint in late 2018.
What started as an exploration of available data ended up being quite an interesting study and we learned a lot from it. We got quite a lot of positive social media attention, and I even won the best talk prize at PopGroup52! We knew many people cared, so we submitted the manuscript to a fancy journal with a broad readership and indeed, the manuscript was sent out for a review. So far so good…
However the reviewer comments, although welcoming our effort of jointly interpreting all the genomic evidence available, concluded that our results were not novel or surprising. Getting burned once was not enough. The next submission was still aiming high, but we ended up with disturbingly similar comments.
The thing is, plenty of the original genome papers suggested some consequences of parthenogenesis. Our reanalysis of 26 genomes revealed that most of them if not all were either lineage-specific, or, at best, a lineage-specific consequence of parthenogenesis. However, would you guess where the individual genomes got published? So, I learned as a young aspiring scientist that a single genome (n=1) is a more successful publication strategy than to quantitatively evaluate predictions across a range of taxa. Alright that was maybe too bleak, but to be completely honest, I still feel very weird about it.
Our work kind of confirmed a previously suspected problem – that lineage specific evolution has stronger effects than parthenogenesis itself. Hence, a proper interpretation of the data would require lineage-specific controls. Unfortunately, almost no species analyzed had a sexual sister known or sequenced, therefore it was hard to systematically take the sexual sisters into account. That is, however, something we were able to do in Californian stick insects (check our fresh preprint). As an exclusive content of this blogpost, here is a heterozygosity figure combining measured heterozygosity in the two studies (Figure 1).
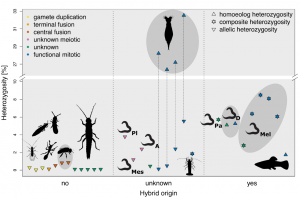
The different submissions and additional analyses took us so long, that by the time we were ready, the long-awaited exciting special issue Sex and Asex: The Genetics of complex life cycles in the Journal of Heredity was about to happen. It was a perfect home for the manuscript alongside other important contributions in our field. And who knows, perhaps in a way the reviews were a blessing in disguise, the special issue in a nice society journal might attract even more attention than in any of general journals.
References



