The first steps in the process of speciation are a bit paradoxical when you think about it…how does one freely interbreeding species make the transition to two reproductively isolated, independent species? More specifically, how do intraspecific mating barriers become interspecific? And why even are there intraspecific mating barriers? Well, that last question is easier to answer, so let’s start there. It starts with inbreeding…more specifically, its prevention.
Inbreeding in general can be a risky move (see: the Hapsburg Dynasty). In flowering plants, inbreeding –specifically selfing – provides reproductive assurance, but also lowers a populations ability to adapt to novel problems. Flowering plants have developed several methods to prevent self-fertilization (floral design, herkogamy, protandry, etc.), but a genetic mechanism that has been well studied is the S-locus and its role in self pollen recognition. This was first described in the familySolanaceaeand species can essentially be put into two groups; self-compatible (SC) and self-incompatible (SI). Simply stated, for a SI species, imagine a population with four S alleles (S0, S1, S2, S3). A female plant with the genotype S0S1(heterozygous for different S alleles) can only accept pollen carrying the S2or S3haplotype. Pollen carrying the S0or S1haplotypes will be rejected as that may be pollen from the same plant. The S-locus encodes for a protein (S-RNase) that is expressed in the style and degrades the developing pollen tube ~1/3 of the way down the style, killing the pollen grain in the process. This type of intraspecific mating barrier would be classified as “postmating prezygotic” isolation. It occurs afterthe pollination event, but beforethe pollen reaches the ovule.
Another example of a postmating prezygotic barrier is unilateral incompatibility (UI). UI occurs when pollen is rejected in “only one direction of an interspecific cross” (Jewell et al. 2020). UI follows a general rule of thumb that a SI species can reject heterospecific pollen (pistil-side rejection) that comes from a SC species, but notpollen from a different (heterospecific) SI species. Conversely, SC species are unable to reject pollen from any parent, implying that pollen has the ability to “neutralize” pistil-side rejection (hereafter referred to as “pollen-side” avoidance), but this has been lost in some SC species (Bedinger et al. 2017). For this reason, it is thought that the genetic mechanisms underlying UI may be linked to those controlling SI.
In the wild tomato clade (Solanum sect. Lycopersicum), there is intraspecific variation in the strength of UI response. Within a particular species, populations that have recently shifted from SI to SC show a slowed or complete loss of interspecific UI response (Baek et al. 2015). This intraspecific variation afforded Jewell and their co-authors an opportunity to uncover whether or not the gene(s) that control UI are the same (or similar) to those that control SI pistil-side rejection.
Using two lines of Solanum pennellii, one SC (LA0716) and one SI (LA3778), Jewell et al. (2020) conducted a QTL (Quantitative Trait Locus) study to answer this question. A QTL study allows researchers to find a statistical association between a phenotype of interest (Quantitative Trait) and a genotype at a particular locus (Locus). By taking two parents differing in some phenotype (in this case, UI response) and crossing them, you get an F1 that is heterozygous at every marker in the genome (in this case, 569 markers – read below). By letting the F1’s mate, recombination shuffles up the entire genome and you can begin to look for patterns.
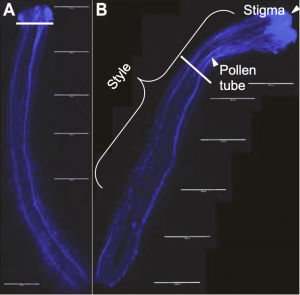
In addition to differing in self-compatibility, these lines also differ in their UI response with the SI line showing a much more rapid response. They crossed the two lines, using the SI LA3778 as the pollen donor to create F1’s, which were self-compatible, and were then selfed to make an F2 mapping population (n=100) to measure strength of UI. Each F2 was then emasculated and pollinated with pollen from S. lycopersicum (LA3475, SC), a common pollen donor as it elicits a “highly repeatable” UI response (Baek et al. 2015). One day after pollination, styles were collected and pollen tubes were visualized (Figure 1). UI was measured by taking the average length of the 5 longest pollen tubes divided by the total length of the style. The same measurements were taken for the parental lines and the F1’s.
The UI response in the F2’s spanned the distribution of both parent lines (proportion of style length in F2’s = 0.01 – 0.55, SI parent = 0.038 ± 0.005, SC parent = 0.32 ± 0.16). These UI scores were then compared to the genotype at particular locations in the genome in order to find an association (the proverbial bread and butter of this paper).
To uncover the genetic basis, Jewell and their colleagues (2020) identified 569 genetic markers that were fixed between the two parent genotypes (e.g. hypothetical marker #362: A/A in SI parent, G/G in SC parent) to follow through to the F2’s. These markers identified a single large effect QTL on Chromosome 1 that explains over 30% of the variation in UI seen in the F2’s. In other words, if a particular individual was G/G at marker #326 that plant was much more likely to have a slower UI response than plants that were A/A or A/G. Additionally, this QTL peak not only overlaps with QTL peaks for UI in other Solanumspecies (indicating a somewhat conserved pathway), but also many of the loci involved with the S-locus system! This finding lends credence to the hypothesis that the underlying mechanism for this interspecific mating barriers (UI) isassociated with the intraspecific mechanism (SI vs. SC).
As the “cherry on top”, the researchers were able to show functional evidence that theS-locus is involved with UI by measuring S-RNase expression in the F2’s. The SC parent is known to not express S-RNase (Li and Chetelat 2015), so researchers hypothesized that the variation in UI response may be associated with S-RNase expression. A subset of the F2’s representing the slowest (n=12) and quickest (n=9) UI response were also assayed for protein expression, and those that expressed the protein had much more rapid UI responses, further implicating the S-locus in inter- and intraspecific pollen rejection.
Now, how does all of this intraspecific variation in UI relate to speciation? Well, it is worth noting that the SC genotype had a slower UI response – in other words, it wasn’t as good at rejecting the wrong pollen – and this appears to be aided by a non-functional S-RNase protein. This loss can lead to a gain of “pistil-side” UI for the SI lineage. When the SC lineage loses S-RNase functionality (aka loses pistil-side rejection), it also relaxes selection in that population to maintain pollen-side avoidance (Bedinger et al. 2017). When the pollen loses function in the SC lineage, the SI lineage, by default, gains UI rejection against the SC line. Over time this prezygotic barrier may lead to a stronger, more permanent barrier to gene flow. Understanding the development and evolution of this intraspecific variation in Solanumgives us a great opportunity to start to uncover the processes that drive genetic mating barriers and speciation.
Lastly, you are probably wondering at this point…how does hockey come into all of this? Hockey tracks a special statistic in addition to the regular assist – the “hockey assist”. A “hockey assist” is the pass that leads to the pass that leads to the goal (the assist to the assist). Well, loss of S-RNase expression (pass #1) can lead to loss of pollen side functionality (pass #2), which results in a stronger intraspecific mating barrier. Over time, this intraspecific mating barrier might turn into an interspecific mating barrier (goal) after being coupled with other genetic incompatibilities, floral shape transitions, or phenological shifts. Loss of S-RNase is the pass that leads to the pass that leads to the goal (speciation).
And understanding that…well that’s the real goal, isn’t it?
References

Zac Cabin is a 6thyear PhD candidate at University of California, Santa Barbara in Dr. Scott Hodges’ lab. Zac is interested in combining field work and molecular techniques to help understand the selective pressures that cause changes in floral shape and structure and specifically, the molecular signatures of those changes. He is currently working on a rare, naturally occurring mutant of Aquilegia coeruleaand the affect of non-pollinator agents of selection on flower shape. Follow Zac @zcabin.



