(feature image: The wild sunflower growing by the roadside. Credit: Matt Lavin from Plants of the World Online. Facilitated by the Royal Botanic Gardens, Kew.)
Sabrina Heiser (@SabrinaHeiser) has written blog posts in Dr. Stacy Krueger-Hadfield’s Conservation Genetics and Science Communication courses at the University of Alabama at Birmingham. She is passionate about science communication, and, therefore, takes any opportunity she can to advance her skills. Her current research focuses on the evolutionary processes generating and maintaining diversity in a red macroalgal species in Antarctica. You can read more about the research she has been involved in on UAB in Antarctica or get a glimpse of what else she is up to on her website.
We’ve been moving organisms around the world for a very long time. While we may have only intended introductions for cultivation purposes, preventing unintentional spread is next to impossible.
Hernández et al. (2019) set out to unravel the invasion history of Helianthus annuus, which has been moved around for cultivation purposes in order to harvest them for their oil and seeds.
Another commonly cultivated sunflower, H. petiolaris, hybridizes with H. annuus, leading to hybrid zones both within the native and non-native ranges.
The wild sunflower originated in the central part of North America. Around 500 years ago, it expanded its range towards the south as well as the west of the present-day US. Later it was introduced to Argentina and to southern Australia.
In Argentina, the introduction is thought to have occurred ~70 years ago, but we do not know much about its invasion history. Hernández et al. (2019) used nuclear and chloroplast simple sequence repeat (SSR) markers, as well as phenotypic variables, to trace the journey of the wild sunflower.
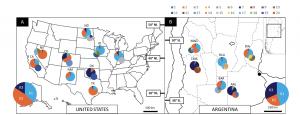
The central and western US were dominated by single haplotypes respectively, whereas the southern US populations had a mix of the most dominant haplotypes (Hernández et al. 2019, Figure 1). All of these haplotypes (and some more) were also present in Argentina indicating multiple introductions.
The three US regions were also differentiated by phenotypic characteristics. The Argentinian populations overlapped with all three of the US regions, with the exception of one population that clustered on its own.
In order to unravel the timing and complexity of the invasion, the authors used ABC (Figure 2).
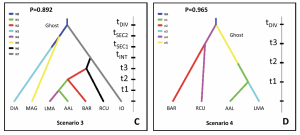
More sampling of more locations within the native and introduced ranges is required in order to further unravel the exact origin of populations in Argentina – although, which study would not benefit from more sampling! It might even be necessary to sample locations from Europe as potential additional sources for genetic and phenotypic variation. Furthermore, hybridization with local species, admixture, and genetic drift also contributed to the tangled web of invasion in Argentina.
Stay tuned for future studies to unravel further “Where did you come from, where did you go? Where did you come from Helianthus annuusL.?”
References





